This is the fourth of four posts on the Corded Ware—Uralic identification:
- Corded Ware—Uralic (I): Differences and similarities with Yamna
- Corded Ware—Uralic (II): Finno-Permic and the expansion of N-L392/Siberian ancestry
- Corded Ware—Uralic (III): “Siberian ancestry” and Ugric-Samoyedic expansions
- Corded Ware—Uralic (IV): Haplogroups R1a and N in Finno-Ugric and Samoyedic
Let me begin this final post on the Corded Ware—Uralic connection with an assertion that should be obvious to everyone involved in ethnolinguistic identification of prehistoric populations but, for one reason or another, is usually forgotten. In the words of David Reich, in Who We Are and How We Got Here (2018):
Human history is full of dead ends, and we should not expect the people who lived in any one place in the past to be the direct ancestors of those who live there today.
Haplogroup N
Another recurrent argument – apart from “Siberian ancestry” – for the location of the Uralic homeland is “haplogroup N”. This is as serious as saying “haplogroup R1” to refer to Indo-European migrations, but let’s explore this possibility anyway:
Ancient haplogroups
We have now a better idea of how many ancient migrations (previously hypothesized to be associated with westward Uralic migrations) look like in genetic terms. From Damgaard et al. (Science 2018):
These serial changes in the Baikal populations are reflected in Y-chromosome lineages (Fig. SA; figs. S24 to S27, and tables S13 and SI4). MAI carries the R haplogroup, whereas the majority of Baikal_EN males belong to N lineages, which were widely distributed across Northern Eurasia (29), and the Baikal_LNBA males all carry Q haplogroups, as do most of the Okunevo_EMBA as well as some present-day Central Asians and Siberians.
The only N1c1 sample comes from Ust’Ida Late Neolithic, 180km to the north of Lake Baikal, which – together with the Bronze Age sample from the Kola peninsula, and the medieval sample from Ust’Ida – gives a good idea of the overall expansion of N subclades and Siberian ancestry among the Circum-Arctic peoples of Eurasia, speakers of Palaeo-Siberian languages.
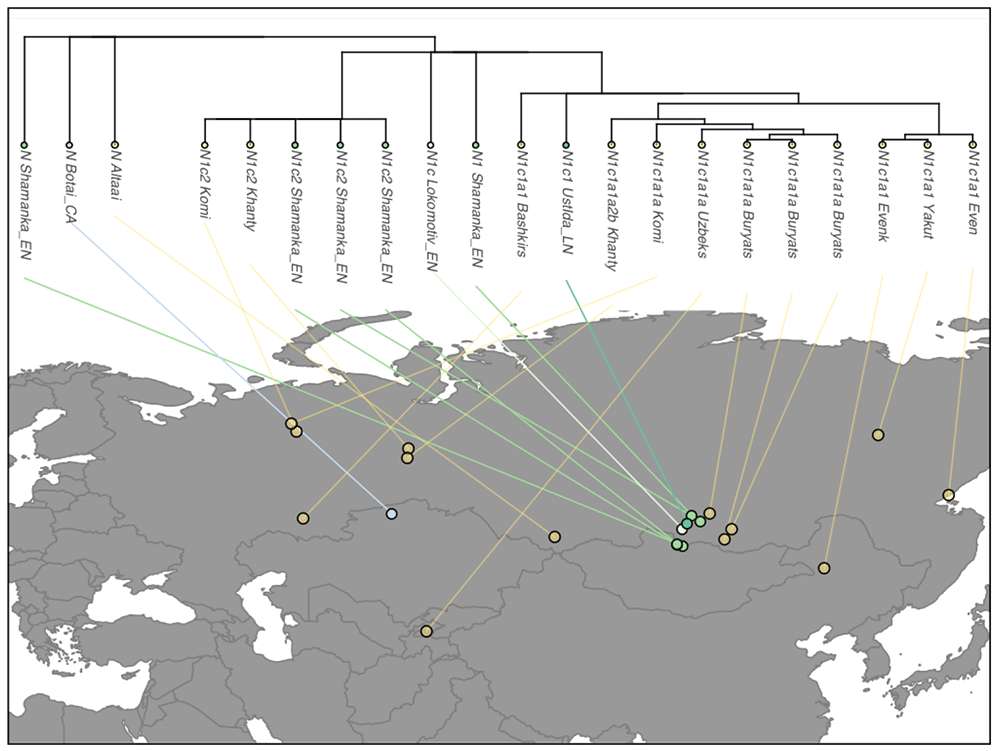
Modern haplogroups
What we should expect from Uralic peoples expanding with haplogroup N – seeing how Yamna expands with R1b-L23, and Corded Ware expands with R1a-Z645 – is to find a common subclade spreading with Uralic populations. Let’s see if it works like that for any N-X subclade, in data from Ilumäe et al. (2016):
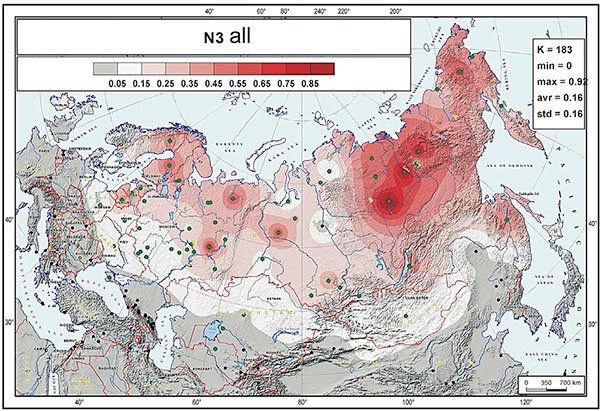
Within the Eurasian circum-Arctic spread zone, N3 and N2a reveal a well-structured spread pattern where individual sub-clades show very different distributions:
N1a1-M46 (or N-TAT), formed ca. 13900 BC, TMRCA 9800 BC
N1a1a2-B187, formed ca. 9800 BC, TMRCA 1050 AD:
The sub-clade N3b-B187 is specific to southern Siberia and Mongolia, whereas N3a-L708 is spread widely in other regions of northern Eurasia.
N1a1a1a-L708, formed ca. 6800 BC, TMRCA 5400 BC.
N1a1a1a2-B211/Y9022, formed ca. 5400 BC, TMRCA 1900 BC:
The deepest clade within N3a is N3a1-B211, mostly present in the Volga-Uralic region and western Siberian Khanty and Mansi populations.
N1a1a1a1a-L392/L1026), formed ca. 4400 BC, TMRCA 2800 BC:
The neighbor clade, N3a3’6-CTS6967, spreads from eastern Siberia to the eastern part of Fennoscandia and the Baltic States
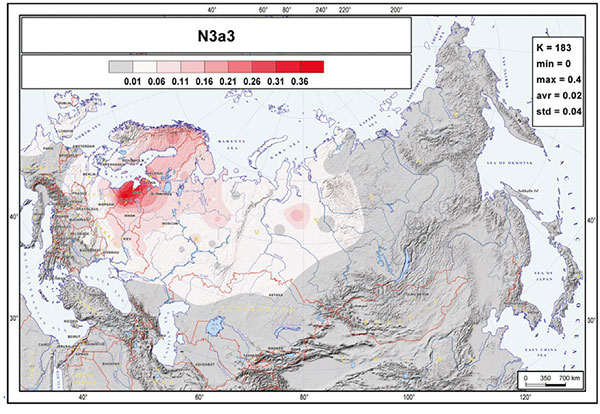
N1a1a1a1a1a-CTS2929/VL29, formed ca. 2100 BC, TMRCA 1600 BC:
In Europe, the clade N3a3-VL29 encompasses over a third of the present-day male Estonians, Latvians, and Lithuanians but is also present among Saami, Karelians, and Finns (Table S2 and Figure 3). Among the Slavic-speaking Belarusians, Ukrainians, and Russians, about three-fourths of their hg N3 Y chromosomes belong to hg N3a3.
In the post on Finno-Permic expansions, I depicted what seems to me the most likely way of infiltration of N1c-L392 lineages with Akozino warrior-traders into the western Finno-Ugric populations, with an origin around the Barents sea.
This includes the potential spread of (a minority of) N1c-B211 subclades due to contacts with Anonino on both sides of the Urals, through a northern route of forest and forest-steppe regions (equivalent to the distribution of Cherkaskul compared to Andronovo), given the spread of certain subclades in Ugric populations.
NOTE. An alternative possibility is the association of certain B211 subclades with a southern route of expansion with Pre-Scythian and Scythian populations, under whose influence the Ananino culture emerged -which would imply a very quick infiltration of certain groups of haplogroup N everywhere among Finno-Ugrics on both sides of the Urals – , and also the expansion of some subclades with Turkic-speaking peoples, who apparently expanded with alliances of different peoples. Both (Scythian and Turkic) populations expanded from East Asia, where haplogroup N (including N1c) was present since the Neolithic. I find this a worse model of expansion for upper clades, but – given the YFull estimates and the presence of this haplogroup among Turkic peoples – it is a possibility for many subclades.
N1a1a1a1a2-Z1936, formed ca. 2800 BC, TMRCA 2400 BC:
The only notable exception from the pattern are Russians from northern regions of European Russia, where, in turn, about two-thirds of the hg N3 Y chromosomes belong to the hg N3a4-Z1936—the second west Eurasian clade. Thus, according to the frequency distribution of this clade, these Northern Russians fit better among other non-Slavic populations from northeastern Europe. N3a4 tends to increase in frequency toward the northeastern European regions but is also somewhat unexpectedly a dominant hg N3 lineage among most Turcic-speaking Volga Tatars and South-Ural Bashkirs.
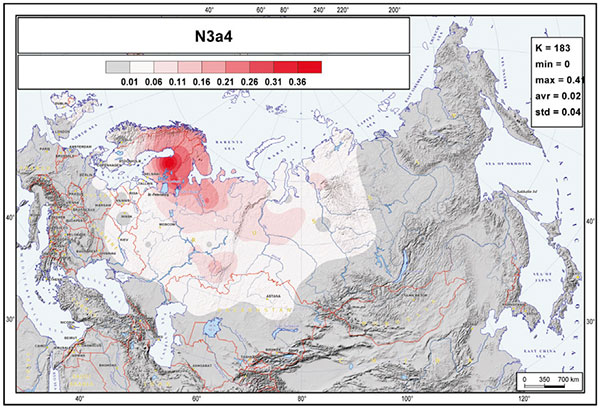
The expansion of N1a-Z1936 in Fennoscandia is most likely associated with the expansion of Saami into asbestos ware-related territory (like the Lovozero culture) during the Late Iron Age – and mixture with its population – , and with the later Fennic expansion to the east and north, replacing their language, as well as with Arctic and forest populations assimilated during Permic, Ugric, and Samoyedic expansions to the north.
N1a1a1a1a4-M2019 (previously N3a2), formed ca. 4400 BC, TMRCA 1700 BC:
Sub-hg N3a2-M2118 is one of the two main bifurcating branches in the nested cladistic structure of N3a2’6-M2110. It is predominantly found in populations inhabiting present-day Yakutia (Republic of Sakha) in central Siberia and at lower frequencies in the Khanty and Mansi populations, which exhibit a distinct Y-STR pattern (Table S7) potentially intrinsic to an additional clade inside the sub-hg N3a2
The second widespread sub-clade of hg N is N2a. (…):
N1a2b-P43 (B523/FGC10846/Y3184), formed ca. 6800 BC, TMRCA ca. 2700 BC:
The absolute majority of N2a individuals belong to the second sub-clade, N2a1-B523, which diversified about 4.7 kya (95% CI = 4.0–5.5 kya). Its distribution covers the western and southern parts of Siberia, the Taimyr Peninsula, and the Volga-Uralic region with frequencies ranging from from 10% to 30% and does not extend to eastern Siberia (…)
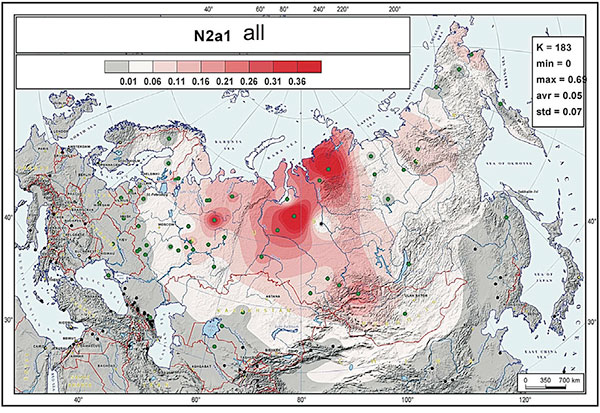
The “European” branch suggested earlier from Y-STR patterns turned out to consist of two clades
N1a2b2a-Y3185/FGC10847, formed ca. 2200 BC, TMRCA 800 BC:
N2a1-L1419, spread mainly in the northern part of that region.
N1a2b2b1-B528/Y24382, formed ca. 900 BC, TMRCA ca. 900 BC:
N2a1-B528, spread in the southern Volga-Uralic region.
Haplogroup R1a
We also have a good idea of the distribution of haplogroup R1a-Z645 in ancient samples. Its subclades were associated with the Corded Ware expansion, and some of them fit quite well the early expansion of Finno-Permic, Ugric, and Samoyedic peoples to the east.
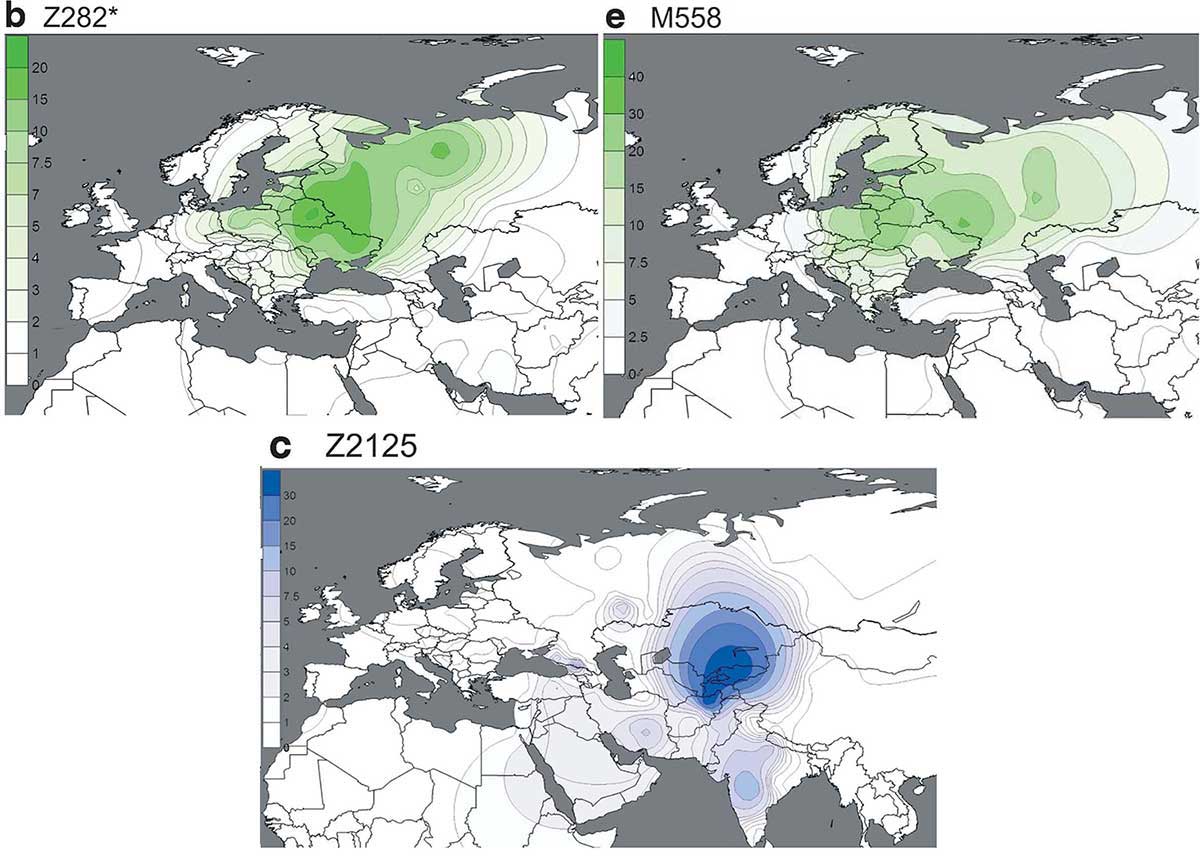
This is how the modern distribution of R1a among Uralians looks like, from the latest report in Tambets et al. (2018):
- Among Fennic populations, Estonians and Karelians (ca. 1.1 million) have not suffered the greatest bottleneck of Finns (ca. 6-7 million), and show thus a greater proportion of R1a-Z280 than N1c subclades, which points to the original situation of Fennic peoples before their expansion. To trust Finnish Y-DNA to derive conclusions about the Uralic populations is as useful as relying on the Basque Y-DNA for the language spread by R1b-P312…
- Among Volga-Finnic populations, Mordovians (the closest to the original Uralic cluster, see above) show a majority of R1a lineages (27%).
- Hungarians (ca. 13-15 million) represent the majority of Ugric (and Finno-Ugric) peoples. They are mainly R1a-Z280, also R1a-Z2123, have little N1c, and lack Siberian ancestry, and represent thus the most likely original situation of Ugric peoples in 4th century AD (read more on Avars and Hungarians).
- Among Samoyedic peoples, the Selkup, the southernmost ones and latest to expand – that is, those not heavily admixed with Siberian populations – , also have a majority of R1a-Z2123 lineages (see also here for the original Samoyedic haplogroups to the south).
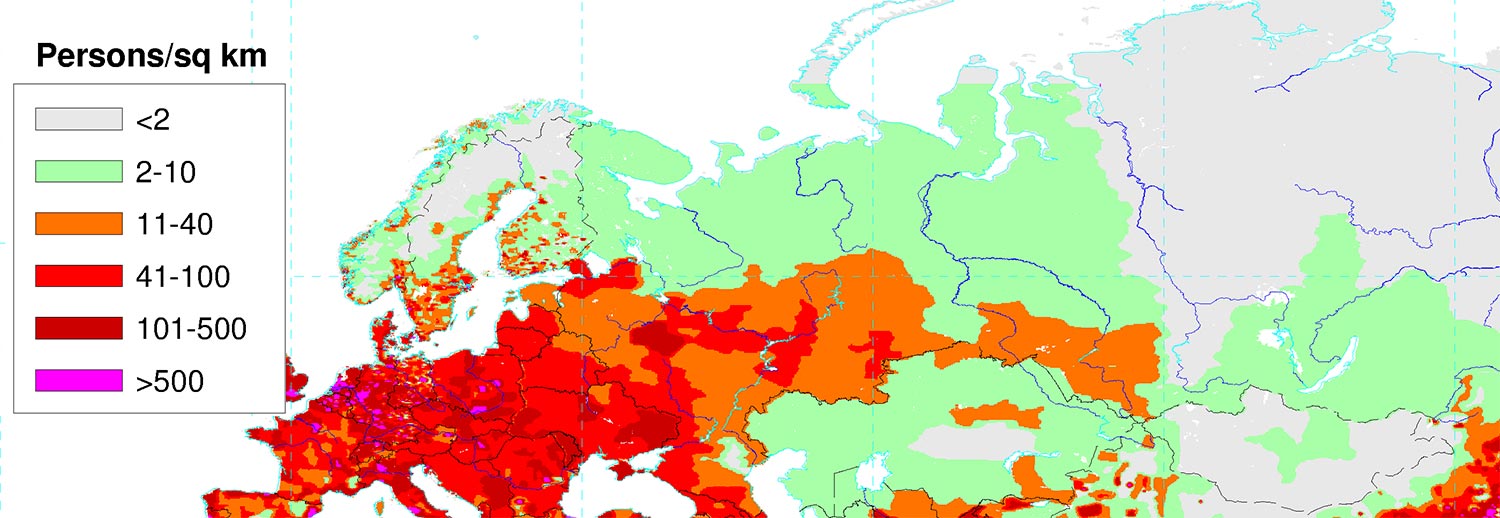
To understand the relevance of Hungarians for Ugric peoples, as well as Estonians, Karelians, and Mordovians (and northern Russians, Finno-Ugric peoples recently Russified) for Finno-Permic peoples, as opposed to the Circum-Arctic and East Siberian populations, one has to put demographics in perspective. Even a modern map can show the relevance of certain territories in the past:
Summary of ancestry + haplogroups
Fennic and Samic populations seem to be clearly influenced by Palaeo-Laplandic peoples, whereas Volga-Finnic and especially Permic populations may have received gene flow from both, but essentially Palaeo-Siberian influence from the north and east.
The fact that modern Mansis and Khantys offer the highest variation in N1a subclades, and some of the highest “Siberian ancestry” among non-Nganasans, should have raised a red flag long ago. The fact that Hungarians – supposedly stemming from a source population similar to Mansis – do not offer the same amount of N subclades or Siberian ancestry (not even close), and offer instead more R1a, in common with Estonians (among Finno-Samic peoples) and Mordvins (among Volga-Finnic peoples) should have raised a still bigger red flag. The fact that Nganasans – the model for Siberian ancestry – show completely different N1a2b-P43 lineages should have been a huge genetic red line (on top of the anthropological one) to regard them as the Uralian-type population.
We know now that ethnolinguistic groups have usually expanded with massive (usually male-biased) migrations, and that neighbouring locals often ‘resurge’ later without changing the language. That is seen in Europe after the spread of Bell Beakers, with the increase of previous ancestry and lineages in Scandinavia during the formation of the Nordic ethnolinguistic community; in Central-West Europe, with the resurgence of Neolithic ancestry (and lineages) during the Bronze Age over steppe ancestry; and in Central-East Europe (with Unetice or East European Bronze Age groups like Mierzanowice, Trzciniec, or Lusatian) showing an increase in steppe ancestry (and resurge of R1a subclades); none of them represented a radical ethnolinguistic change.
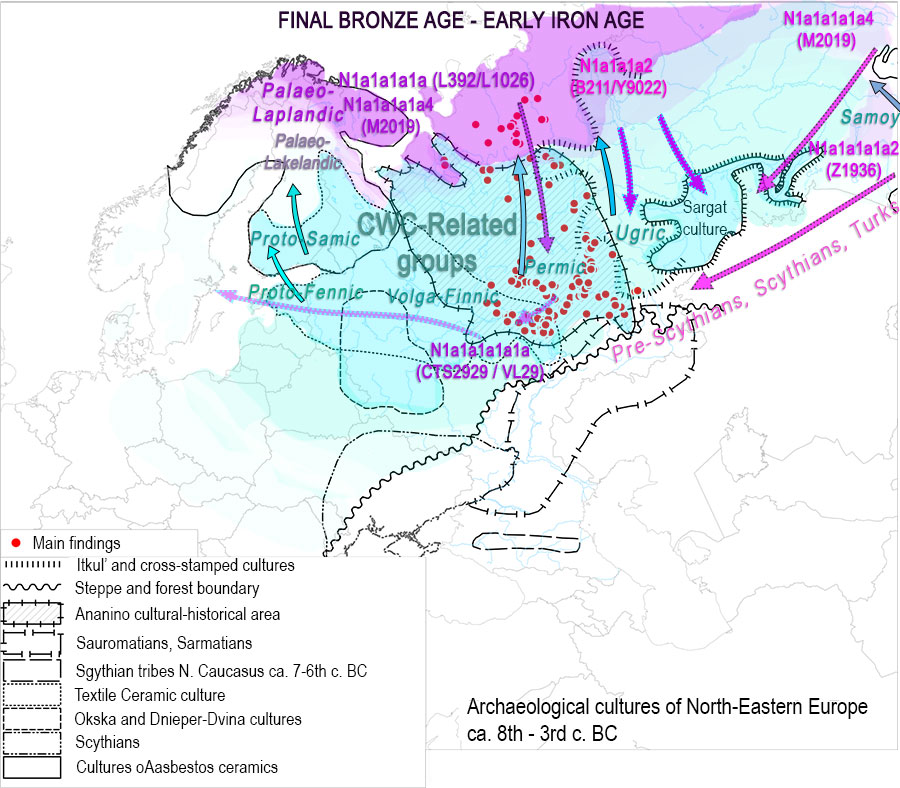
It is not hard to model the stepped arrival, infiltration, and/or resurge of N subclades and “Siberian ancestries”, as well as their gradual expansion in certain regions, associated with certain migrations first – such as the expansions to the Circum-Arctic region, and later the Scythian- and Turkic-related movements – , as well as limited regional developments, like the known bottleneck in Finns, or the clear late expansion of Ugric and Samoyedic languages to the north among nomadic Palaeo-Siberians due to traditions of exogamy and multilingualism. This fits quite well with the different arrival of N (N1c and xN1c) lineages to the different Uralic-speaking groups, and to the stepped appearance of “Siberian ancestry” in the different regions.
The aternative
It is evident that a lot of people were too attached to the idea of Palaeolithic R1b lineages ‘native’ to western Europe speaking Basque languages; of R1a lineages speaking Indo-European and spreading with Yamna; and N lineages ‘native’ to north-eastern Europe and speaking Uralic, and this is causing widespread weeping and gnashing of teeth (instead of the joy of discovering where one’s true patrilineal ancestors come from, and what language they spoke in each given period, which is the supposed objective of genetic genealogy…)
Since an Indo-Germanic branch (as revived now by some in the Copenhaguen group to fit Kristiansen’s theory of the 1980s with recent genetic data) does not make any sense in linguistics, the finding of R1a in Yamna would not have led where some think it would have, because North-West Indo-European would still be the main Late PIE branch in Europe. Don’t take my word for it; take James P. Mallory’s (2013).
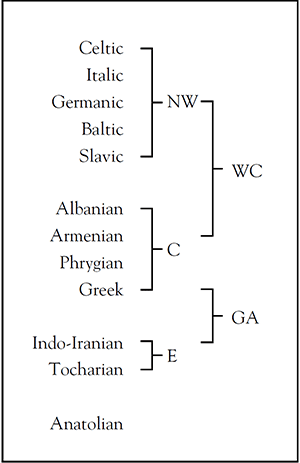
If an (unlikely) Indo-Slavonic group were posited, though, such a group would still be bound (with Indo-Iranian) to the steppes with East Yamna/Poltavka (admixing with Abashevo migrants, but retaining its language), developing Sintashta/Potapovka → Srubna/Andronovo, and R1a lineages would have equally undergone the known bottlenecks of the steppes where they replaced R1b-Z2103 – which this eastern group shares with Balkan languages, a haplogroup that links therefore together the Graeco-Aryan group.
As far as I know – and there might be many other similar pet theories out there – there have been proposals of “modern Balto-Slavic-like” populations (in an obvious circular reasoning based on modern populations) in some Scythian clusters of the Iron Age.
NOTE. I will not enter into “Balto-Slavic-like R1a” of the Late Bronze Age or earlier because no one can seriously believe at this point of development of Population Genetics that autosomal similarity predating 1,500+ years the appearance of Slavs equates to their (ethnolinguistic) ancestral population, without a clear intermediate cultural and genetic trail – something we lack today in the Slavic case even for the late Roman period…
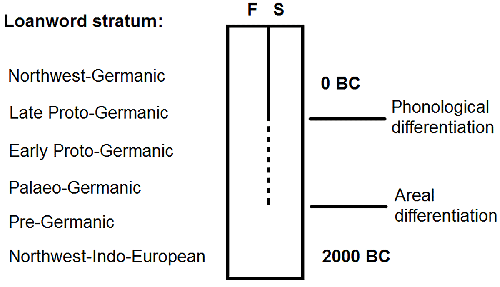
We also know of R1a-Z280 lineages in Srubna, probably expanding to the west. With that in mind, and knowing that Palaeo-Germanic was in close contact with Finno-Samic while both were already separated but still in contact, and that Palaeo-Germanic was also in contact and closely related to a ‘Temematic’ distinct from Balto-Slavic (and also that early Proto-Baltic and Proto-Slavic from the Roman Iron Age and later were in contact with western Uralic) this will be the linguistic map of the Iron Age if R1a is considered to expand Indo-European from some kind of “patron-client” relationship with west Yamna:
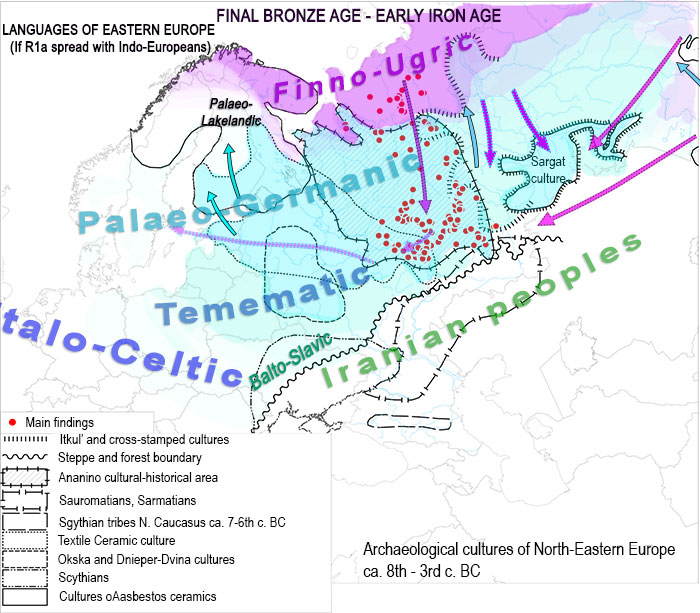
You might think I have some personal or political reason against this kind of proposals. I haven’t. We have been proposing Indo-European to be the language of the European Union for more than 10 years, so to support R1b-Italo-Celtic in the whole Western Europe, R1a-Germanic in Central and Eastern Europe, and R1a-Indo-Slavonic in the steppes (as the Danish group seems to be doing) has nothing inherently bad (or good) for me. If anything, it gives more reason to support the revival of North-West Indo-European in Europe.
My problem with this proposal is that it is obviously beholden to the notion of the uninterrupted cultural, historic and ethnic continuity in certain territories. This bias is common in historiography (von Falkenhausen 1993), but it extends even more easily into the lesser known prehistory of any territory, and now more than ever some people feel the need to corrupt (pre)history based on their own haplogroups (or the majority haplogroups of their modern countries). However, more than on philosophical grounds, my rejection is based on facts: this picture is not what the combination of linguistic, archaeological, and genetic data shows. Period.
Nevertheless, if Yamna + Corded Ware represented the “big and early expansion” of Germanic and Italo-Celtic peoples proper of the dream Nazi’s Lebensraum and Fascist’s spazio vitale proposals; Uralians were Siberian hunter-gatherers that controlled the whole eastern and northern Russia, and miraculously managed to push (ethnolinguistically) Neolithic agropastoralists to the west during and after the Iron Age, with gradual (and often minimal) genetic impact; and Balto-Slavic peoples were represented by horse riders from Pokrovka/Srubna, hiding then somewhere around the forest-steppe until after the Scythian expansion, and then spreading their language (without much genetic impact) during the early Middle Ages…so be it.
See also
- Uralic speakers formed clines of Corded Ware ancestry with WHG:ANE
- Magyar tribes brought R1a-Z645, I2a-L621, and N1a-L392(xB197) lineages to the Carpathian Basin
- R1a-Z280 and R1a-Z93 shared by ancient Finno-Ugric populations; N1c-Tat expanded with Micro-Altaic
Related
- The traditional multilingualism of Siberian populations
- Iron Age bottleneck of the Proto-Fennic population in Estonia
- Y-DNA haplogroups of Tuvinian tribes show little effect of the Mongol expansion
- Corded Ware—Uralic (I): Differences and similarities with Yamna
- Haplogroup R1a and CWC ancestry predominate in Fennic, Ugric, and Samoyedic groups
- The Iron Age expansion of Southern Siberian groups and ancestry with Scythians
- Evolution of Steppe, Neolithic, and Siberian ancestry in Eurasia (ISBA 8, 19th Sep)
- Mitogenomes from Avar nomadic elite show Inner Asian origin
- On the origin and spread of haplogroup R1a-Z645 from eastern Europe
- Oldest N1c1a1a-L392 samples and Siberian ancestry in Bronze Age Fennoscandia
- Consequences of Damgaard et al. 2018 (III): Proto-Finno-Ugric & Proto-Indo-Iranian in the North Caspian region
- The concept of “Outlier” in Human Ancestry (III): Late Neolithic samples from the Baltic region and origins of the Corded Ware culture
- Genetic prehistory of the Baltic Sea region and Y-DNA: Corded Ware and R1a-Z645, Bronze Age and N1c
- More evidence on the recent arrival of haplogroup N and gradual replacement of R1a lineages in North-Eastern Europe
- Another hint at the role of Corded Ware peoples in spreading Uralic languages into north-eastern Europe, found in mtDNA analysis of the Finnish population
- New Ukraine Eneolithic sample from late Sredni Stog, near homeland of the Corded Ware culture