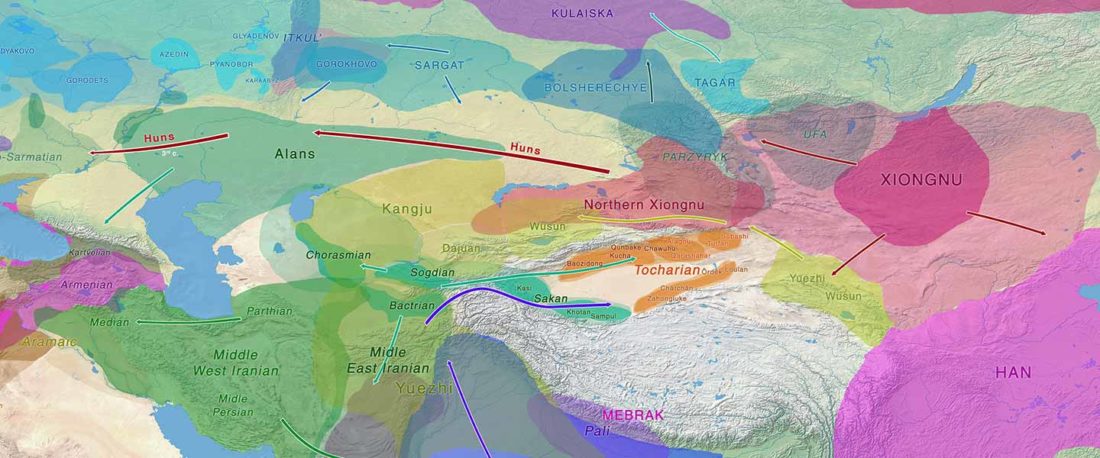
This post is part of a draft on palaeolinguistics and the Proto-Uralic homeland. See below for the color code of protoforms.
14. Earliest PU ~ PIE contacts
14.1. Indo-Uralic?
The most reliable correspondences to propose an Indo-Uralic phylum come from basic morphological comparisons. Some of the most frequently mentioned ones include (e.g. Čop 1975, Kortlandt 2002, Bjørn 2019, or Lubotsky 2019):
- Nominal endings:
- PU nom.sg. *-Ø ~ PIA nom.-acc.sg. *-Ø (in neuter athematic nouns).
- PU acc.sg. *-m ~ PIA acc.sg. *-m.
- PU dual *-ki(-) ~ PIA nom.-acc.du. *-h₁.
- PU abl. *-tA ~