The following are updated files for unsupervised ADMIXTURE of most available ancient Eurasian samples with K=7. For reference, see PCA of ancient and modern Eurasian samples.
NOTE. For a precise interpretation of ancestry evolution, be sure to first check the posts on the expansion of “Steppe ancestry”, on the spread of Yamnaya ancestry with Indo-Europeans, and on the evolution of Corded Ware ancestry typical of modern Uralic populations.
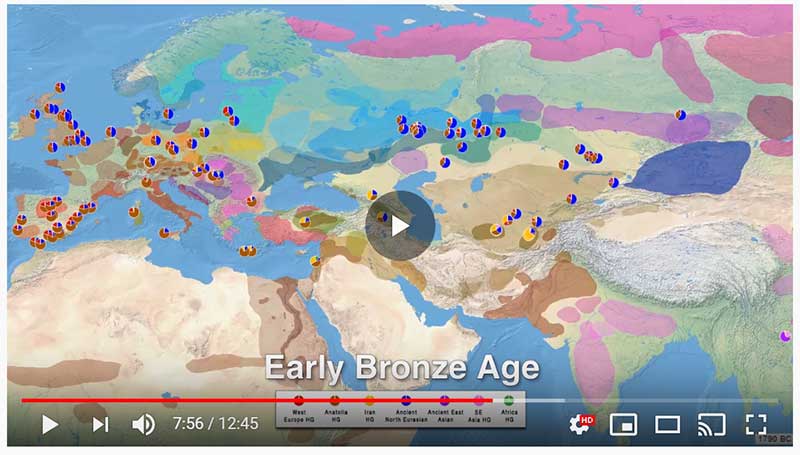
ADMIXTURE timeline
This is a YouTube video similar to the one on Indo-Europeans and Y-DNA evolution:
Some comments
- I have tried running supervised ADMIXTURE models by selecting distant populations based on PCAs and qpAdm results. The most accurate approximations to what the software should offer appear with a small K number, between K=5 and K=7, whether supervised or unsupervised, and adding more ancestral populations gives some weird results the more distant (in time) populations are from these selected samples.
- Labels for ancestral components are used following those commonly referred to in the literature, although supervised ADMIXTURE using corresponding available samples (viz. Anatolia Neolithic for AHG, Iran Hotu and/or CHG for IHG, AG2, AG3 and Mal’ta for ANE, etc.) offer slightly different, less smooth outputs for some periods, especially among more recent populations.
- Outputs depend on many different factors, and these files are intended as an overview of the evolution of these simplistic components. The number of available samples per period, the potential ancestry changes within each conventionally selected period, or whether or not each available sample is representative of the territory they were recovered from, among many other factors, influence the outputs and the maps.
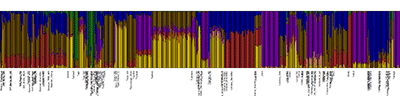
NOTE. In summary, ADMIXTURE results like these below might be used to develop new ideas, to be then formally tested; they cannot be used to support anything. Don’t be like the Copenhagen group, randomly selecting “Steppe ancestry” with K=4, identifying this component as “Indo-Europeans”, and correlating its evolution with changes in vegetation composition in yet another obvious correlation = causation argument among many confounding factors left unaccounted for…
Static ADMIXTURE + culture maps
Colours correspond to the components as labelled in the video and in the files below.
- Anatomically Modern Humans (PDF)
- Upper Palaeolithic (PDF)
- Epipalaeolithic (PDF)
- Early Mesolithic (PDF)
- Late Mesolithic (PDF)
- Neolithic and hunter-gatherer pottery (PDF)
- Early Eneolithic (PDF)
- Late Eneolithic (PDF)
- Early Chalcolithic (PDF)
- Late Chalcolithic (PDF)
- Early Bronze Age (PDF)
- Middle Bronze Age (PDF)
- Late Bronze Age (PDF)
- Early Iron Age (PDF)
- Late Iron Age (PDF)
- Antiquity (PDF)
- Middle Ages (PDF)
Natural interpolation maps of ADMIXTURE
The following maps offer natural neighbour interpolations of ancestral components in ancient DNA samples grouped by periods (conventionally selected following the same pattern as in the Prehistory Atlas).
- Extrapolation (inferred ancestry beyond the frame created by available samples per map) is obtained by adding distant external locations (such as Greenland, Arctic, Alaska…) with a value of 0.
- Videos offer a dynamic timeline.
- Click on the images to see a version with higher resolution.
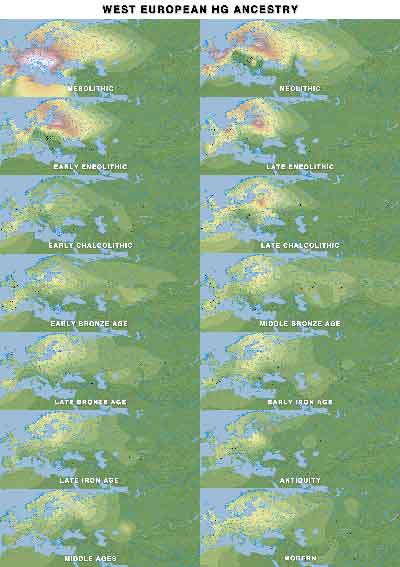
WHG ancestry
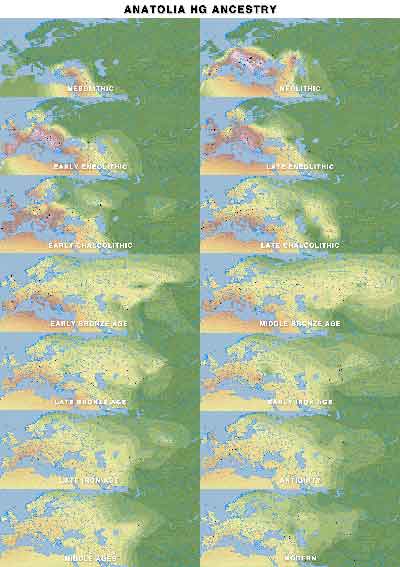
AHG ancestry
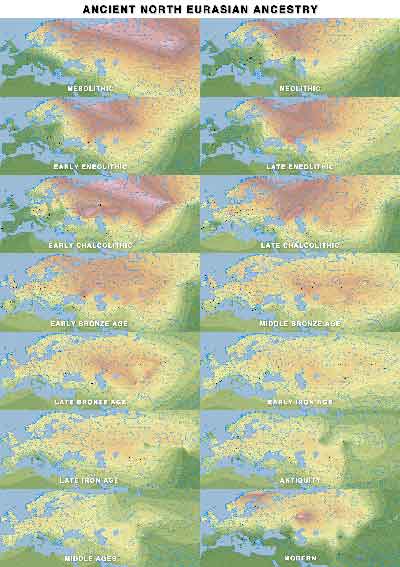
ANE ancestry
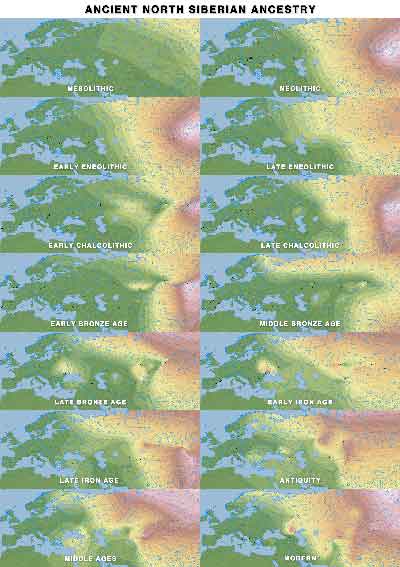
“Siberian” ancestry
This ancestry peaks among Baikal HG, Ust’Belaya, Nganasans, or Ulchi, hence the different labels used.
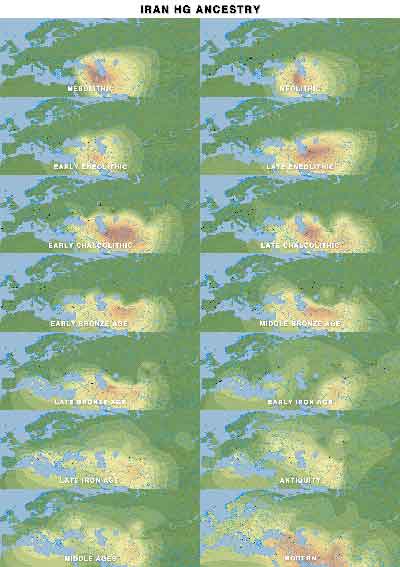
Iran HG ancestry
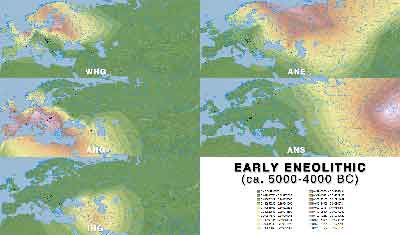
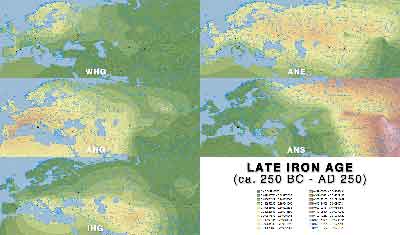
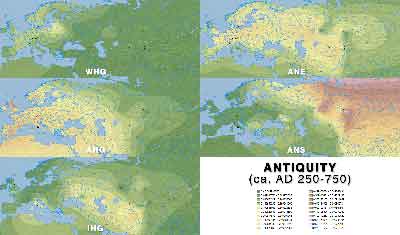
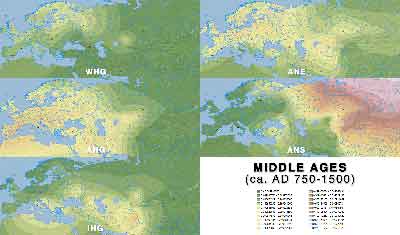
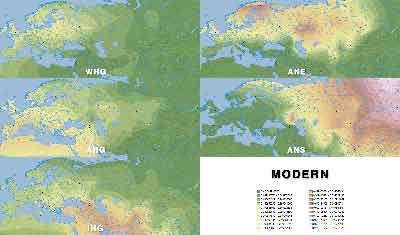
ADMIXTURE maps by period
Click on each image for a higher resolution version.
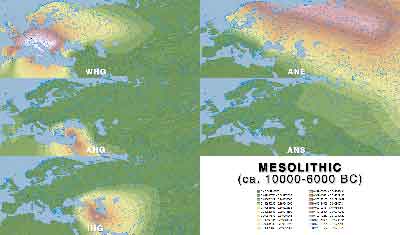
Mesolithic
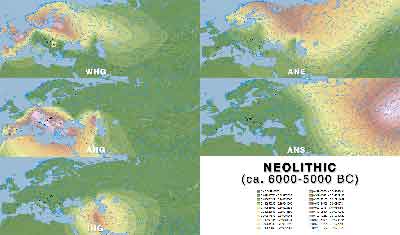
Neolithic
Early Eneolithic
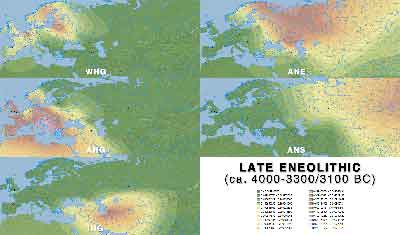
Late Eneolithic
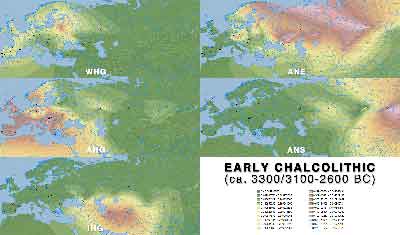
Early Chalcolithic
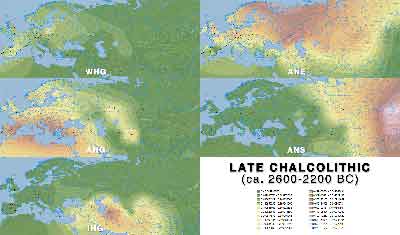
Late Chalcolithic
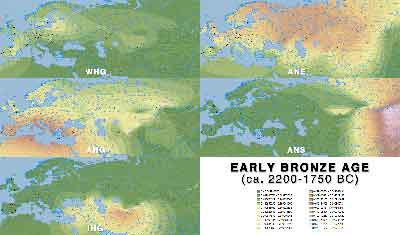
Early Bronze Age
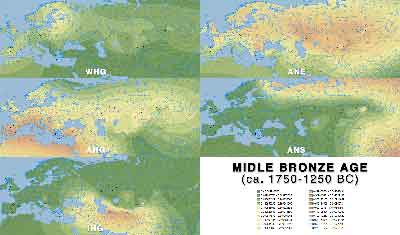
Middle Bronze Age
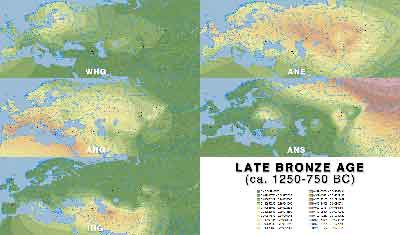
Late Bronze Age
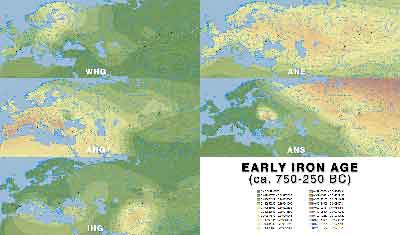
Early Iron Age
Late Iron Age
Antiquity
Middle Ages
Modern populations
Samples
These are the samples used for interpolations in each period (except for modern populations, which are those included in the Reich Lab curated dataset):
- Mesolithic (PDF)
- Neolithic (PDF)
- Early Eneolithic (PDF)
- Late Eneolithic (PDF)
- Early Chalcolithic (PDF)
- Late Chalcolithic (PDF)
- Early Bronze Age (PDF)
- Middle Bronze Age (PDF)
- Late Bronze Age (PDF)
- Early Iron Age (PDF)
- Late Iron Age (PDF)
- Antiquity (PDF)
- Middle Ages (PDF)