New paper (behind paywall) Demographic and Genetic Portraits of the Ulchi Population, by Balanovska et al. Russian Journal of Genetics (2018) 54(10):1245–1253.
Interesting excerpts (emphasis mine):
Marital structure. The intensity of interethnic marriages puts the existence of the Ulchi population at risk. The colorful ethnic composition of the Ulchi settlements is reflected in the marriage structure [see featured image]. We found that the proportion of single-ethnic marriages of the Ulchi is on average 51%. The greatest number of such marriages takes place in the village of Bulava. Marriages of Ulchi with Russians are in second place. Marriages with indigenous peoples of the Far East, Nanais, Nivkhs, Evenks, and others, are in third place. Thus, almost half of the Ulchi marriages are with representatives of other nationalities. Such a significant level of interethnic mixing makes it possible to talk about intense processes of assimilation of this indigenous people and puts to the forefront the problem of loss of the unique gene pool of the Ulchi.
Haplogroup C (its branch M48) was genotyped for its five subbranches with markers M86, B470, F13686, B93, and the marker at position 16645386 (GRCh37), which was found by our team for the first time. Variant B93 is rare in the Ulchi, and 14 samples (that is, more than a quarter of the entire gene pool of the Ulchi, Fig. 2) belong to M86 and its subvariants. Therefore, we genotyped STR markers of C-M86 carriers for the Ulchi and neighboring Amur populations and analyzed the relationships of detected haplotypes on the phylogenetic network (Fig. 3, STR haplotypes are available from authors upon request).
(…) On the network, different clusters are associated with different populations: most Mongols belong to F13686, all Evenks of the Amur River region with this haplogroup form a subcluster within F13686, and part of Upper Nanais is the basis of cluster B470.
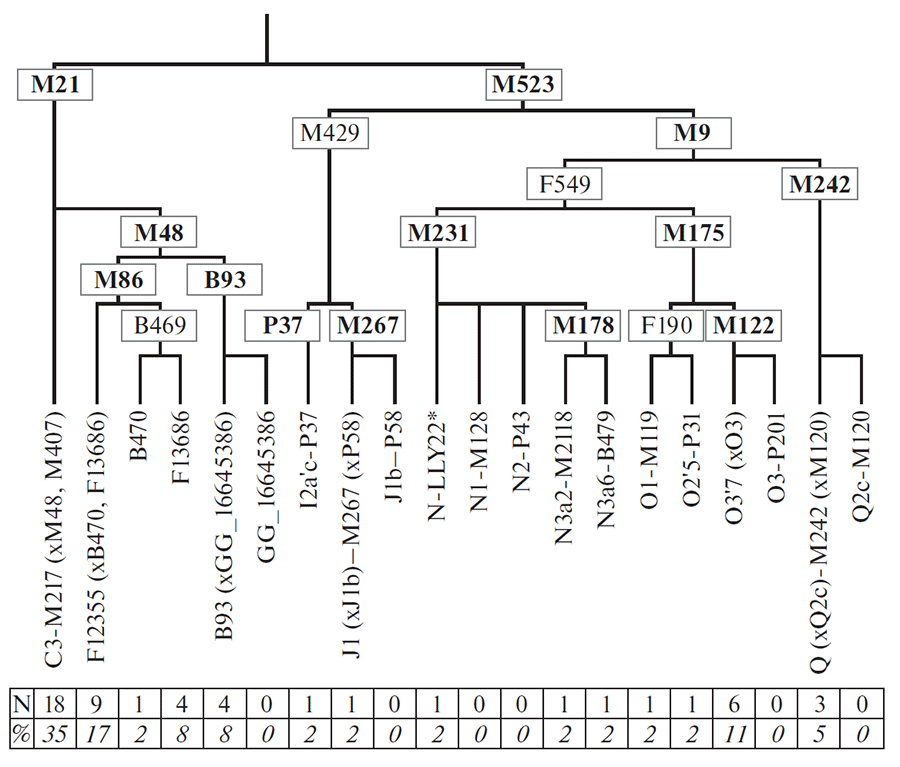
An estimate of the age of the entire haplogroup C-F12355 obtained from the data of genome-wide sequencing of seven specimens is 2400 ± 500 years (O.P. Balanovsky, unpublished data). That is, the common ancestor of all the studied representatives of various peoples with this haplogroup lived not so long ago, the first millennium BC. The formation time of cluster F13686 is somewhat later: 1990 ± 600 years.
(…) obvious traces of the interaction of the gene pool of the Ulchi with neighboring and remote peoples of the Far East and Central Asia in the time range of the last one to three thousand years were revealed. This shows that the results of work [4] on the similarity of the gene pool of the ancient (age of 7500 years) Neolithic genomes of the Amur River region to the Ulchi probably indicate not the uniqueness of the Ulchi, but the fact that this ancient gene pool was preserved in a vast circle of populations of the Far East interwoven with gene flows both with each other and, to a lesser extent, with populations of Central Asia.
The expansion of C2b1a2a-M86 (among many basal C2-M217 samples) is thus possibly associated with the spread of Tungusic, which puts C2b1a at the root of the Micro-Altaic expansion, with a formation date ca. 12700 BC, TMRCA 12500 BC (and not only Mongolian). This shows that Micro-Altaic is connected with a local population which shows a clear continuity since at least 3500 BC. This, however, tells us little about the origin of the language.
See also the recent ISBA presentation on the Houtaomuga site, Neolithic transition in Northeast Asia; and also Bronze Age population dynamics and rise of dairy pastoralism in Mongolia, Impact of colonization in north-eastern Siberia
That leaves the ancestral N lineages found among Far East Asians as Palaeo-Siberian in origin, and their late expansions to the west not particularly linked with any of the known Palaeo-Siberian ethnolinguistic groups, let alone a supposed “Uralo-Altaic” language…
Related
- Y-DNA haplogroups of Tuvinian tribes show little effect of the Mongol expansion
- Mitogenomes from Avar nomadic elite show Inner Asian origin
- Ancient nomadic tribes of the Mongolian steppe dominated by a single paternal lineage
- Model for the spread of Transeurasian (Macro-Altaic) communities with farming
- Y chromosome C2*-star cluster traces back to ordinary Mongols, rather than Genghis Khan
- Expansion of peoples associated with spread of haplogroups: Mongols and C3*-F3918, Arabs and E-M183 (M81)
- Genomic history of Northern Eurasians includes East-West and North-South gradients