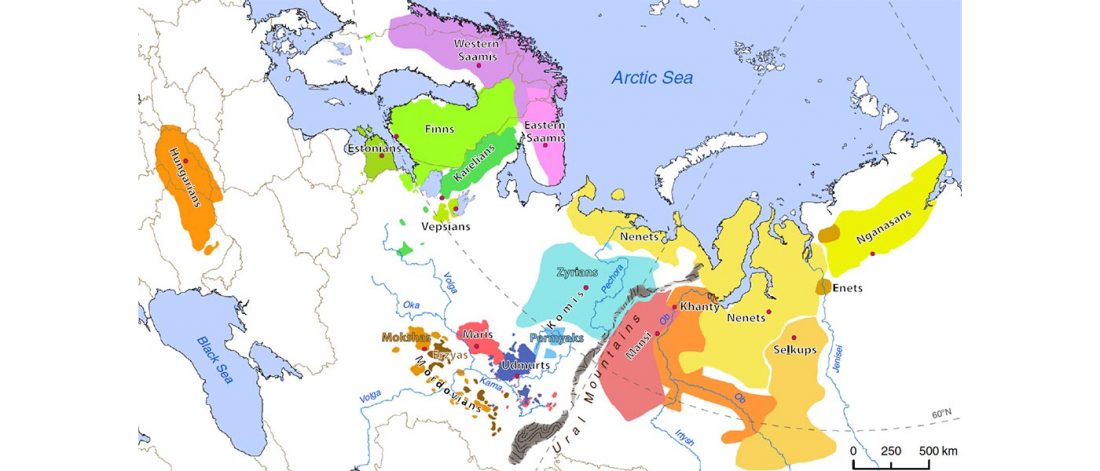
Open access The Arrival of Siberian Ancestry Connecting the Eastern Baltic to Uralic Speakers further East, by Saag et al. Current Biology (2019).
Interesting excerpts:
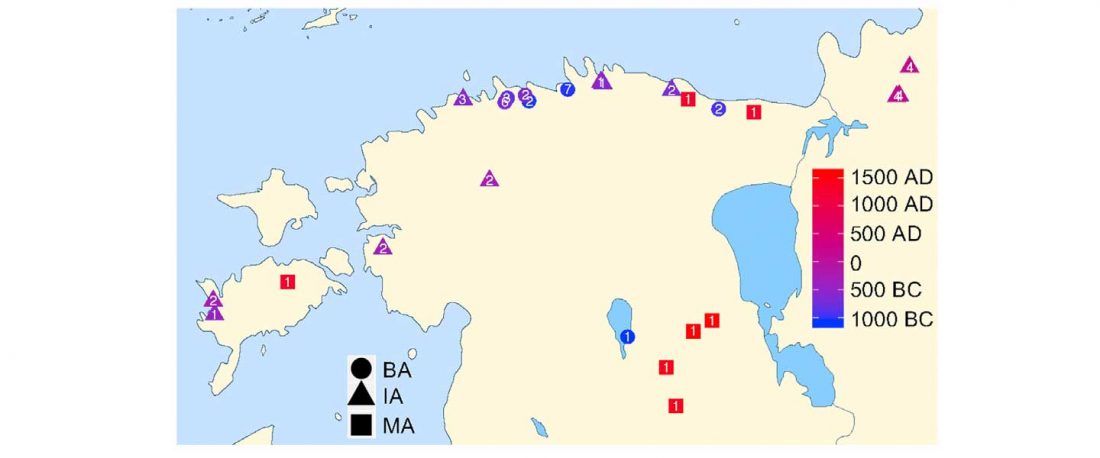
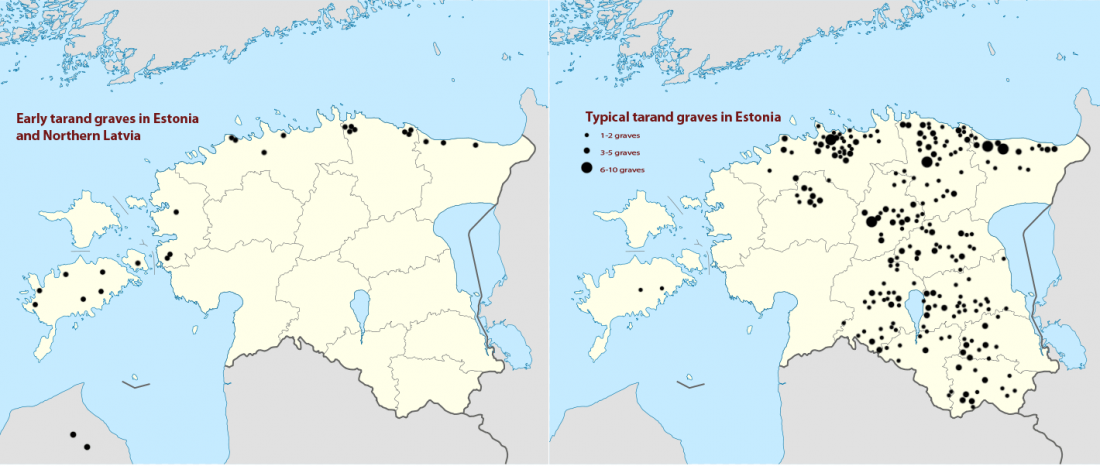
… Read the rest “Baltic Finns in the Bronze Age, of hg. R1a-Z283 and Corded Ware ancestry”In this study, we present new genomic data from Estonian Late Bronze Age stone-cist graves (1200–400 BC) (EstBA) and Pre-Roman Iron Age tarand cemeteries (800/500 BC–50 AD) (EstIA). The cultural background of stone-cist graves indicates strong connections both to the west and the east [20, 21]. The Iron Age (IA) tarands have been proposed to mirror “houses of the dead” found among Uralic peoples of the Volga-Kama region [22].
(…) The 33 individuals included