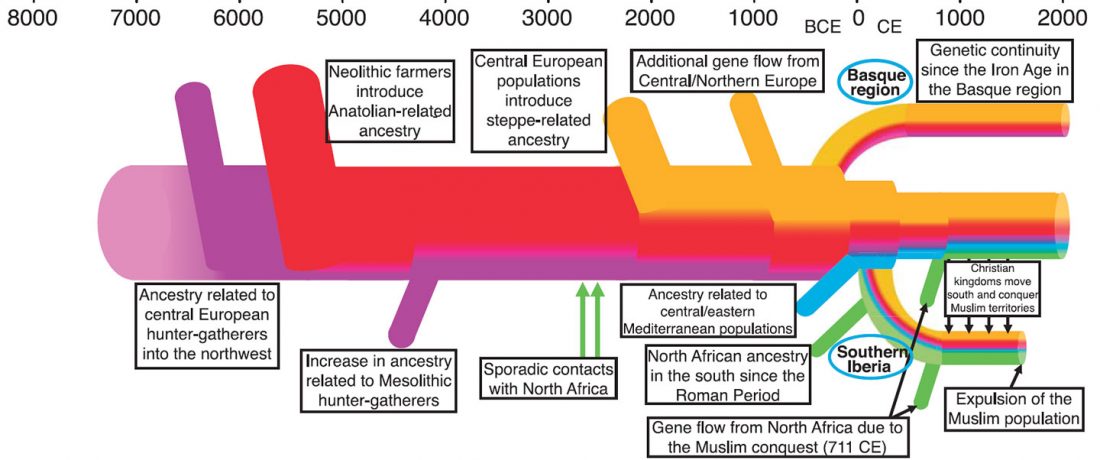
Iberia: East Bell Beakers spread Indo-European languages; Celts expanded later
New paper (behind paywall), The genomic history of the Iberian Peninsula over the past 8000 years, by Olalde et al. Science (2019).
NOTE. Access to article from Reich Lab: main paper and supplementary materials.
Abstract:
… Read the rest “Iberia: East Bell Beakers spread Indo-European languages; Celts expanded later”We assembled genome-wide data from 271 ancient Iberians, of whom 176 are from the largely unsampled period after 2000 BCE, thereby providing a high-resolution time transect of the Iberian Peninsula. We document high genetic substructure between northwestern and southeastern hunter-gatherers before the spread of farming. We reveal sporadic contacts between Iberia and North Africa by ~2500 BCE and, by ~2000 BCE, the replacement