New open access Ancient Genomes Reveal Yamnaya-Related Ancestry and a Potential Source of Indo-European Speakers in Iron Age Tianshan, by Ning et al. Current Biology (2019).
Interesting excerpts (emphasis mine, changes for clarity):
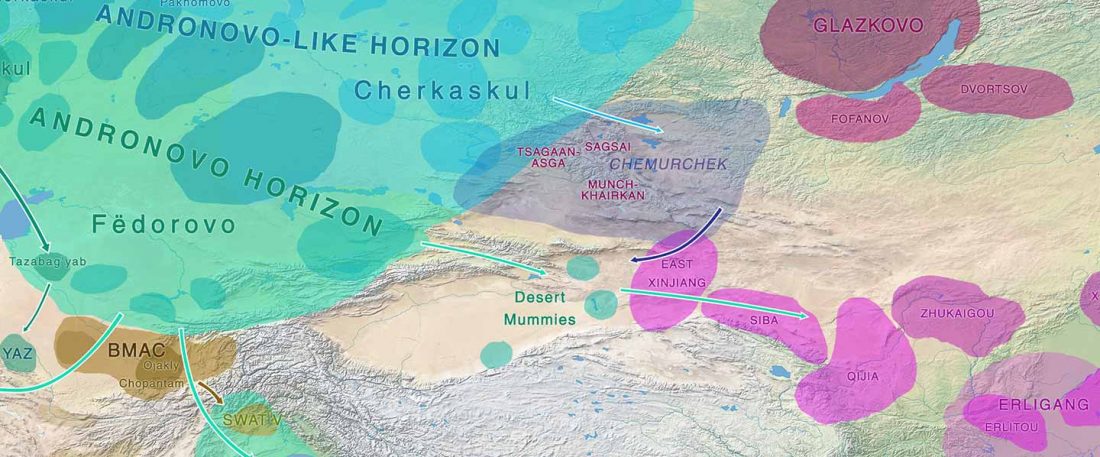
… Read the rest “Iron Age Tocharians of Yamnaya ancestry from Afanasevo show hg. R1b-M269 and Q1a1”Here, we report the first genome-wide data of 10 ancient individuals from northeastern Xinjiang. They are dated to around 2,200 years ago and were found at the Iron Age Shirenzigou site. We find them to be already genetically admixed between Eastern and Western Eurasians. We also find that the majority of the East Eurasian ancestry in the Shirenzigou individuals is related to northeastern Asian populations,