Native American genetic continuity and oldest mtDNA hg A2ah in the Andean region
Native American gene continuity to the modern admixed population from the Colombian Andes: Implication for biomedical, population and forensic studies by Criollo-Rayo et al., Forensic Sci Int Genet (2018), in press, corrected proof.
Abstract (emphasis mine):
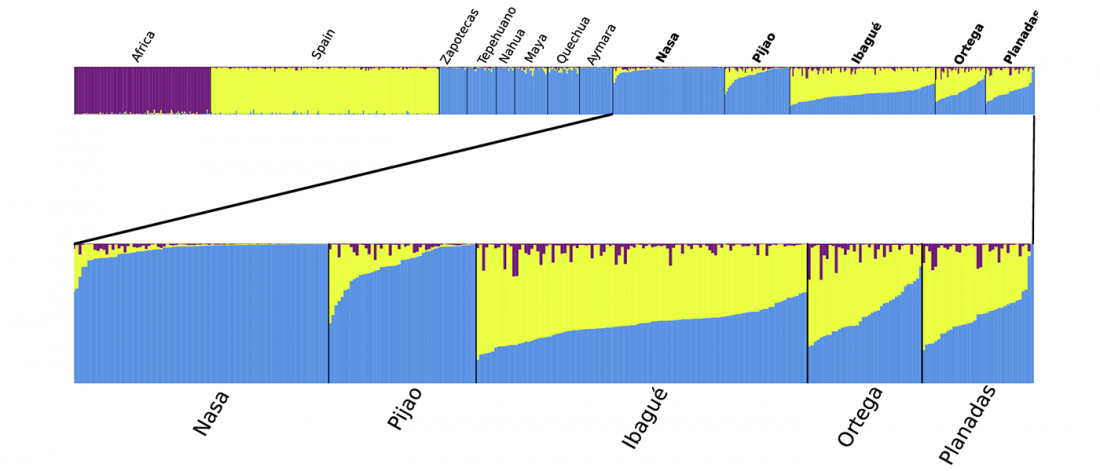
… Read the rest “Native American genetic continuity and oldest mtDNA hg A2ah in the Andean region”Andean populations have variable degrees of Native American and European ancestry, representing an opportunity to study admixture dynamics in the populations from Latin America (also known as Hispanics). We characterized the genetic structure of two indigenous (Nasa and Pijao) and three admixed (Ibagué, Ortega and Planadas) groups from Tolima, in the Colombian Andes. DNA samples from 348 individuals were genotyped for six mitochondrial DNA