Native American gene continuity to the modern admixed population from the Colombian Andes: Implication for biomedical, population and forensic studies by Criollo-Rayo et al., Forensic Sci Int Genet (2018), in press, corrected proof.
Abstract (emphasis mine):
Andean populations have variable degrees of Native American and European ancestry, representing an opportunity to study admixture dynamics in the populations from Latin America (also known as Hispanics). We characterized the genetic structure of two indigenous (Nasa and Pijao) and three admixed (Ibagué, Ortega and Planadas) groups from Tolima, in the Colombian Andes. DNA samples from 348 individuals were genotyped for six mitochondrial DNA (mtDNA), seven non-recombining Y-chromosome (NRY) region and 100 autosomal ancestry informative markers. Nasa and Pijao had a predominant Native American ancestry at the autosomal (92%), maternal (97%) and paternal (70%) level. The admixed groups had a predominant Native American mtDNA ancestry (90%), a substantial frequency of European NRY haplotypes (72%) and similar autosomal contributions from Europeans (51%) and Amerindians (45%). Pijao and nearby Ortega were indistinguishable at the mtDNA and autosomal level, suggesting a genetic continuity between them. Comparisons with multiple Native American populations throughout the Americas revealed that Pijao, had close similarities with Carib-speakers from distant parts of the continent, suggesting an ancient correlation between language and genes. In summary, our study aimed to understand Hispanic patterns of migration, settlement and admixture, supporting an extensive contribution of local Amerindian women to the gene pool of admixed groups and consistent with previous reports of European-male driven admixture in Colombia.
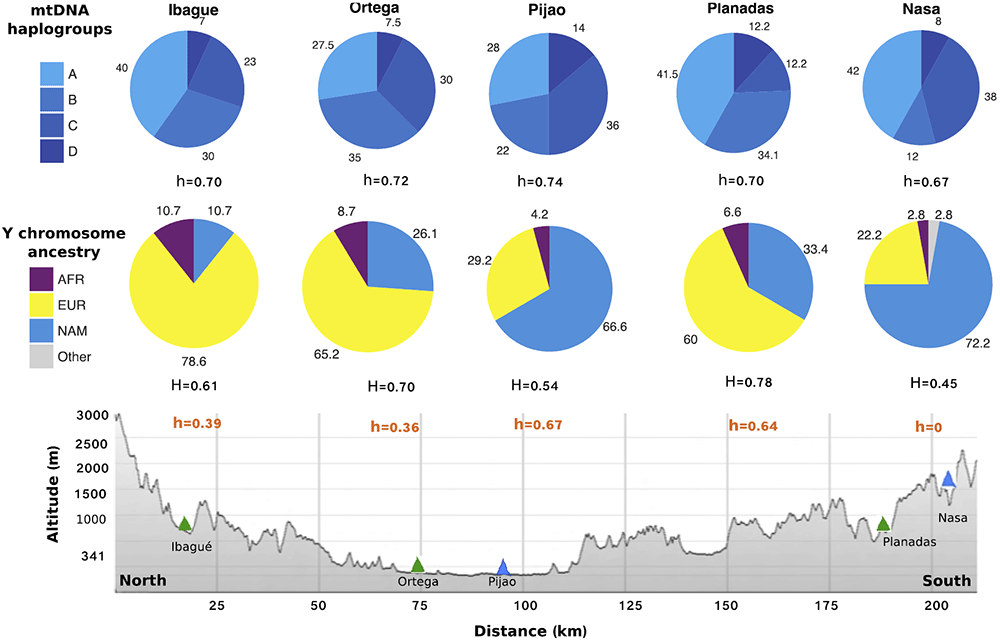
top and middle sections show the frequency of Native American mtDNA haplogroups and NRY lineages for all
populations. Gene diversity is shown below their respective pie chart. The lower part depicts the geography of the
region where the sampling sites of Ortega and Pijao are closely located in Tolima’s Magdalena river valley and
Ibague, Planadas and Nasa located in the Andes cordilleras (additional geographic details are shown in SF1).
Highlights from the paper:
- MtDNA suggest a pre/post Columbian genetic continuity in the Colombian Andes.
- Y-chromosome diversity follows a clinal gradient in the studied region.
- Sex-biased/male-driven admixture process, involving Pijao women with European men.
- Admixed closer to Indigenous resguardos have a higher Native American ancestry.
Also interesting is the recent paper Mitochondrial lineage A2ah found in a pre‐Hispanic individual from the Andean region, by Russo et al., in American Journal of Human Biology (2018), with an interesting sample from the Regional Developments II period (540 ± 60 BP).

Related:
- Male-biased expansions and migrations also observed in Northwestern Amazonia
- Ancient human parallel lineages within North America contributed to a coastal expansion
- Population structure in Argentina shows most European sources of South European origin
- Latin Americans show widespread Mediterranean and North African ancestry
- Patterns of genetic differentiation and the footprints of historical migrations in the Iberian Peninsula
- Ancient Patagonian genomes suggest origin and diversification of late maritime hunter-gatherers
- Ancient DNA reveals temporal population structure of pre-Incan and Incan periods in South‐Central Andes area
- Paternal lineages mainly from migrants, maternal lineages mainly from local populations in Argentina