Recently published paper Genomic Data from an Ancient European Battlefield Indicates On-Going Strong Selection on a Genomic Region Associated with Lactase Persistence Over the Last 3,000 Years, by Burger et al., submitted to Current Biology, available at CellPress SneakPeek.
Interesting excerpts (emphasis mine):
Tollense sample shows no structure
Multiple lines of evidence point to little or no genetic structure in the population from which the Tollense individuals were sampled. First, all individuals fall within the range of Central and northern European variation when projected onto a principle component analysis (PCA) trained on modern samples and their spread matches that of other ancient population samples of similar age; namely the Lech valley in Bavaria (~4,700 to ~3,250 BP; Mittnik et al. 2019,) and Mokrin in Serbia (~4,100 to ~3,700 BP; Žegarac et al. in preparation).
NOTE. For more on the Mokrin cemetery published in Žegarac et al. bioRxiv (2020), see Maros shows Yamnaya-derived East BBC ancestry and local admixture. For more on the Lech Valley samples from Mittnik et al. (2019), see Corded Ware and Bell Beaker related groups defined by patrilocality and female exogamy.

Tollense population is from Central / Northern Europe
The ancestry of the 14 Tollense individuals is Central to northern European, as indicated by several analyses. First, the 70% density contour in the PCA shows a slightly more northern center for the Tollense sample than for those of the Bavarian Lech Valley and Serbian Mokrin individuals. Second, the Tollense sample appears closest to 5th century Bavarians (Veeramah et al. 2018) and modern Central and northern Europeans when quantified with FST, although all European populations have overlapping confidence intervals.
NOTE. For more on Bavarians published in Veeramah et al. (2018), see Genomic analysis of Germanic tribes from Bavaria show North-Central European ancestry.
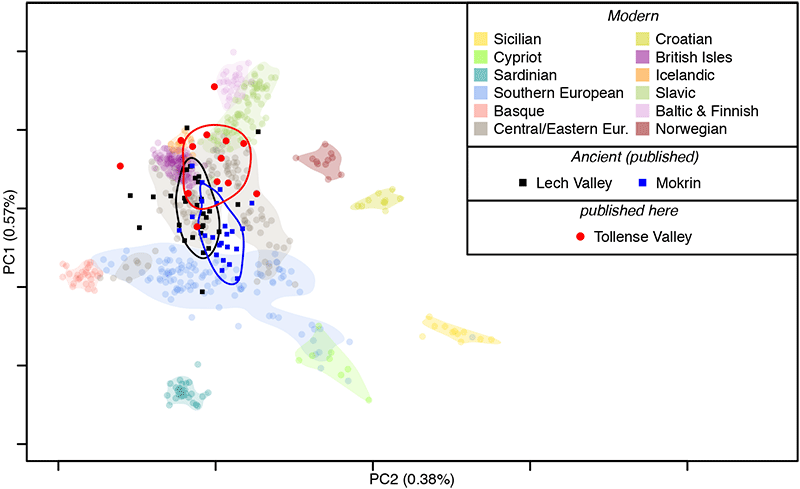
Based on archeological finds at the battlefield, it has been hypothesized that some of the warriors came from southern Central Europe, i.e. south eastern Germany and Bohemia (Uhlig et al. 2019; Terberger et al. 2014; Jantzen 2011). Carbon and Strontium isotope ratios further suggest a mixture of locals and non-locals (Price et al. 2019). While it is possible that our data are insufficient to resolve genetic affinities at such a small geographic scale, it appears likely that the Tollense individuals were sampled from a relatively homogeneous population with a high degree of continuity to those living in the same broad region today.
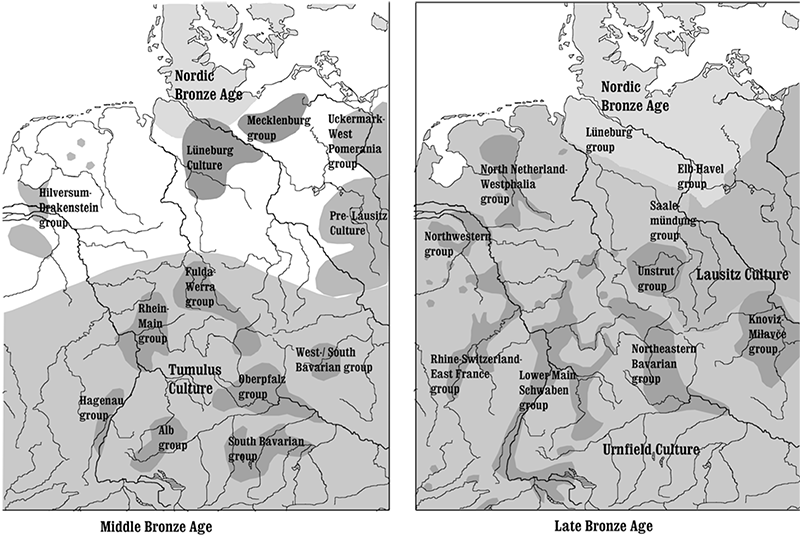
Eastern Europeans steppes are not the source for lactase persistence
Based on imputed data, Allentoft et al. (2015) reported a low allele frequency (5%) of the rs4988235*T allele in Bronze Age Europeans, similar to that reported here, but a much higher frequency (~25%) among Bronze Age samples from the Pontic-Caspian Steppe, indicating a possible steppe origin of lactase persistence. Since imputing allele presence in ancient samples using modern reference individuals may be problematic in regions of strong recent selection, we investigated this hypothesis by genotyping the rs4988235 locus in Eneolithic and Early Bronze Age samples from eastern Europe and the Steppe region. We could not detect the rs4988235*T allele among any of these N=37 samples, suggesting that the frequency of this allele was very low, and possibly 0; almost certainly lower than the 5.4% previously reported for a geographically, culturally and temporally diverse sample with “Steppe ancestry” (Mathieson et al. 2015). While these estimates are not directly informative about the origin of the rs4988235*T allele, they appear inconsistent with a major contribution of the Steppe-associated expansion to the high frequencies observed after the Bronze Age in Europe.
Y-DNA haplogroups
7 out of 16 male samples show haplogroup R1b (one of them hg. R1, likely also R1b), the 3 with best inferences are R1b-P312, 2 others are R1b-L51, the other R1b-M269. The other 9 show haplogroup I2a, probably all I2a-M223, the only one with a more detailed subclade is I2a-Z2059.
#EDIT (16 SEP 2020): The R1 outlier, WEZ56, is of hg. R1a-Z283 (xM458, xZ92). The I2 guys are all likely within the I-Y3672 trunk, and there are one R1b-Z2118*, two R1b-L51 (either L151* or even L51*) and two other R1b-P312.
#EDIT: Ancient DNA Dataset updated and samples uploaded to ArcGIS Online Web App.
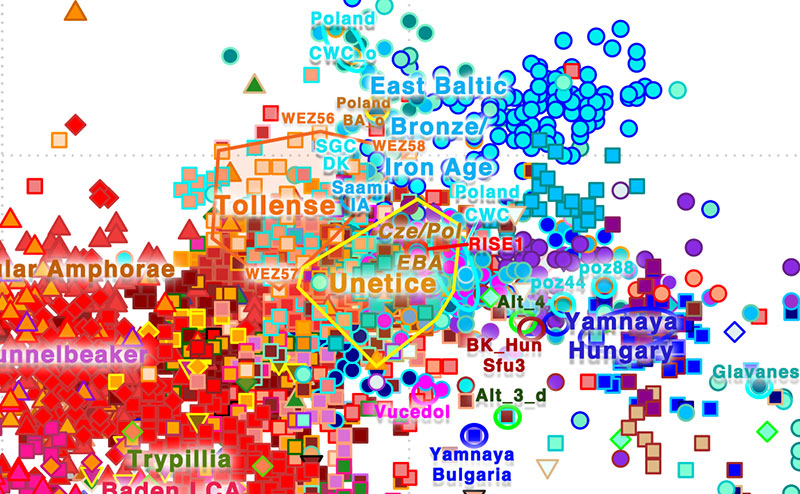
Comments
After an initial massive migration and replacement event during the Bell Beaker expansion, there is an evident admixture between R1b-rich North-West Indo-Europeans and the non-Indo-European-speaking local populations they encountered during the European Early Bronze Age, with further replacement events (including Y-DNA bottlenecks) and acculturation processes obscuring the original palaeolinguistic and archaeogenomic picture.
In this case, it was already known that the Weltzin samples (dated to the transition of the Middle to the Late Bronze Age) show a variation slightly more (W)HG-like than what is found among later Germanic and Early Slavic peoples, which suggests a constant influx of southern ancestry during the Indo-Europeanization of Northern Europe, with pockets of more HG-like peoples surviving up to the Early Middle Ages in Scandinavia.
This tiny sampling from Pomerania was probably made up of a majority of speakers of “Late Pre-Proto-Germanic” or “Early Proto-Germanic” by the Late Bronze Age, based on the expansion of Nordic material culture over a likely Northern Indo-European-speaking population of the Western Baltic. These samples might thus show an ancestry and Y-DNA distribution very similar to the Proto-Balto-Slavic-speaking population that will be sampled in the Lusatian culture.
#EDIT (16 SEP 2020): Complementing the discussion below:
- Supporting the nature of these peoples as directly related to Pre-Proto-Germanic peoples is the fact that subclades R1b-Z2118 and I2-Y3672 (with a quite recent TMRCA just before this battle) are found among Longobards.
- WEZ56 seems to be an outlier in ancestry as in haplogroup, based on the other available ones; however, SNP calls can’t discard that he belongs to an R1a-Y2395 subclade – which was also present among Longobards.
- In fact, strontium analyses show he and WEZ57 (the R1b-Z2118* guy) probably spent their childhood in Scandinavia (Price et al. 2017), fully compatible with the hypothesized origin of Longobards in southern Scandinavia.
Related
- The Lusatian culture, the most likely vector of Balto-Slavic expansions
- European hydrotoponymy (IV): tug of war between Balto-Slavic and West Uralic
- The Tollense Valley battlefield: the North European ‘Trojan war’ that hints to western Balto-Slavic origins
- The significance of the Tollense Valley in Bronze Age North-East Germany
- Consequences of O&M 2018 (III): The Balto-Slavic conundrum in Linguistics, Archaeology, and Genetics
- Tug of war between Balto-Slavic and West Uralic (II)
- European hydrotoponymy (III): from Old European to Palaeo-Germanic and the Nordwestblock
- “Dinaric I2a” and the expansion of Common Slavs from East-Central Europe
- Common Slavs from the Lower Danube, expanding with haplogroup E1b-V13?
- European hydrotoponymy (I): Old European substrate and its relative chronology
- The cradle of Russians, an obvious Finno-Volgaic genetic hotspot
- Aquitanians and Iberians of haplogroup R1b are exactly like Indo-Iranians and Balto-Slavs of haplogroup R1a
- Balto-Slavic accentual mobility: an innovation in contact with Balto-Finnic