Open access FADS1 and the timing of human adaptation to agriculture, by Sara Mathieson & Iain Mathieson, bioRxiv (2018).
Abstract:
Variation at the FADS1/FADS2 gene cluster is functionally associated with differences in lipid metabolism and is often hypothesized to reflect adaptation to an agricultural diet. Here, we test the evidence for this relationship using both modern and ancient DNA data. We document pre-out-of-Africa selection for both the derived and ancestral FADS1 alleles and show that almost all the inhabitants of Europe carried the ancestral allele until the derived allele was introduced approximately 8,500 years ago by Early Neolithic farming populations. However, we also show that it was not under strong selection in these populations. Further, we find that this allele, and other proposed agricultural adaptations including variants at LCT/MCM6, SLC22A4 and NAT2, were not strongly selected until the Bronze Age, 2,000-4,000 years ago. Similarly, increased copy number variation at the salivary amylase gene AMY1 is not linked to the development of agriculture although in this case, the putative adaptation precedes the agricultural transition. Our analysis shows that selection at the FADS locus was not tightly linked to the development of agriculture. Further, it suggests that the strongest signals of recent human adaptation may not have been driven by the agricultural transition but by more recent changes in environment or by increased efficiency of selection due to increases in effective population size.
Interesting excerpt for the steppe-related expansion:
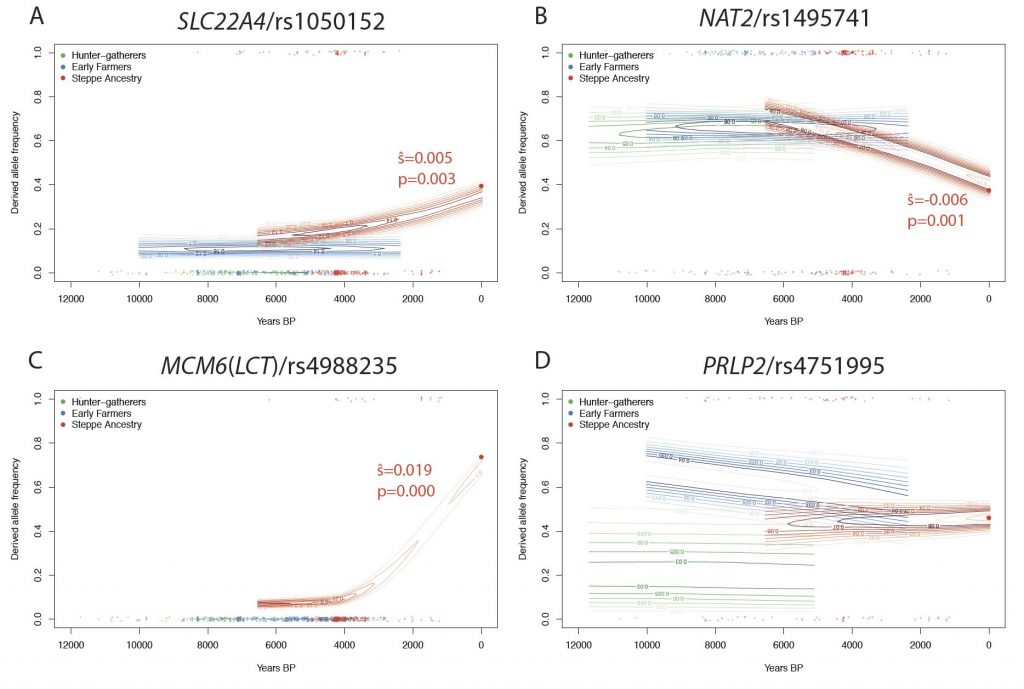
In the case of FADS1 and all the other examples we investigated, the proposed agricultural adaption was either not temporally linked with agriculture or showed no evidence of selection in agricultural populations. Instead, most of the variants with any evidence of selection were only strongly selected at some point between the Bronze Age and the present day, that is, in a period starting 2000-4000 BP and continuing until the present. This time period is one in which there is relatively limited ancient DNA data, and so we are unable to determine the timing of selection any more accurately. Future research should address the question of why this recent time period saw the most rapid changes in apparently diet associated genes. One plausible hypothesis is that the change in environment at this time was actually more dramatic than the earlier change associated with agriculture. Another is that effective population sizes were so small before this time that selection did not operate efficiently on variants with small selection coefficients. For example, analysis of present-day genomes from the United Kingdom suggests that effective population size increased by a factor of 100-1000 in the past 4500 years (Browning and Browning 2015). Ancient effective population sizes less that 104 would suggest that those populations would not be able to efficiently select for variants with selection coefficients on the order of 10-4 or smaller. Larger ancient DNA datasets from the past 4,000 years will likely resolve this question.
This complexity of the reasons for selection reminded me of the comment by Narasimhan on lactase persistence expanding with steppe populations into Central Asia (based on data of the paper where he is the first author):
Examining lactase persistence in the South and Central Asian data, shows that the allele arrives in South Asia with steppe pastoralists, (data from https://t.co/HPE0TOgd5a …) pic.twitter.com/kG66oRg23k
— Vagheesh Narasimhan (@vagheesh) April 18, 2018
I always thought that to argue for natural selection in humans (viz. skin color, lactase persistence, etc.) was possible for archaic groups over tens of thousands of years, but that more recent selections would be very difficult to prove, in so far as historical population expansions involve more ‘artificial’ (i.e. man-made or man-caused) societal changes.
NOTE. I am probably more inclined to think about regional outbreaks (especially of new diseases) as one of the few potential short-term selection mechanisms in historical societies, because of their potential to create sudden bottlenecks of better fitted survivors.
I think recent works like these are showing a mixed situation, where maybe some traits were strongly selected for environmental reasons; but most of the time they were probably – like, say, Y-DNA haplogroup bottlenecks in Europe after the steppe-related expansions – due mostly to chance.