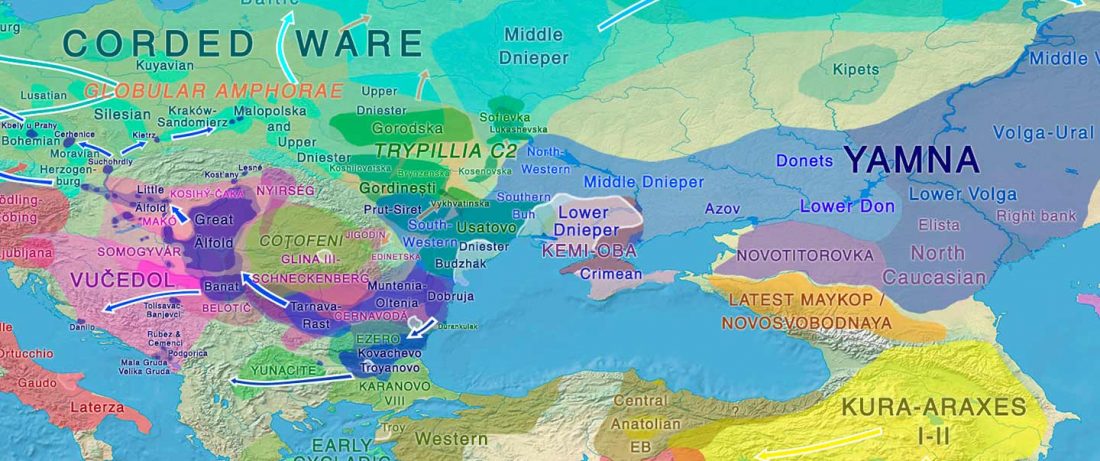
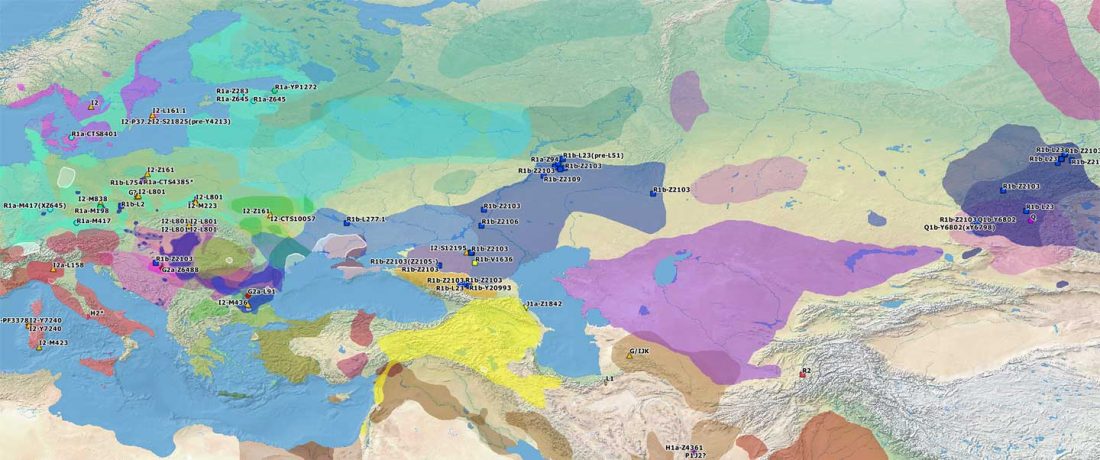
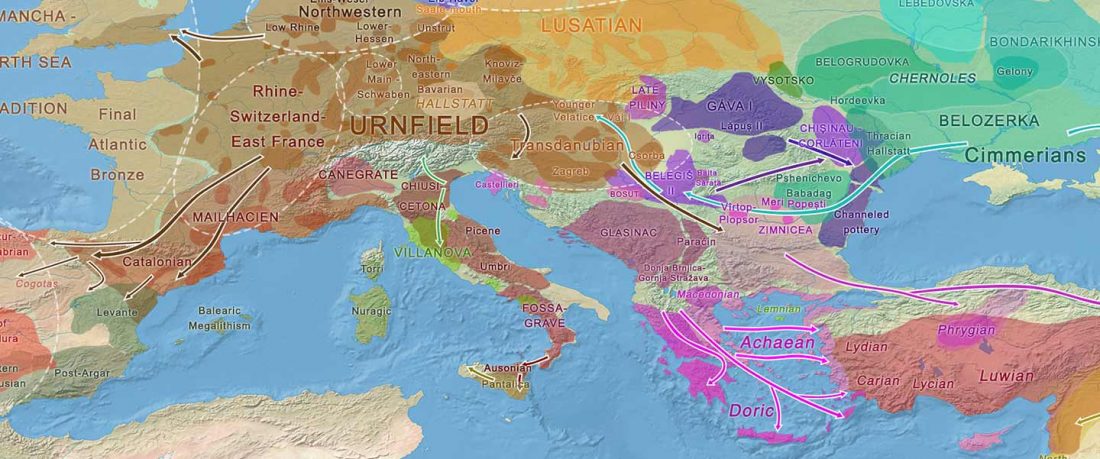
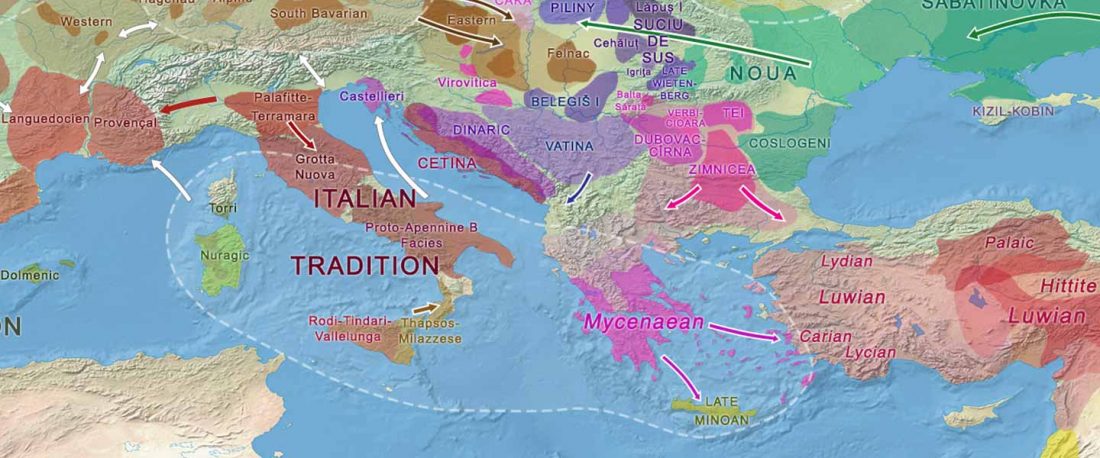
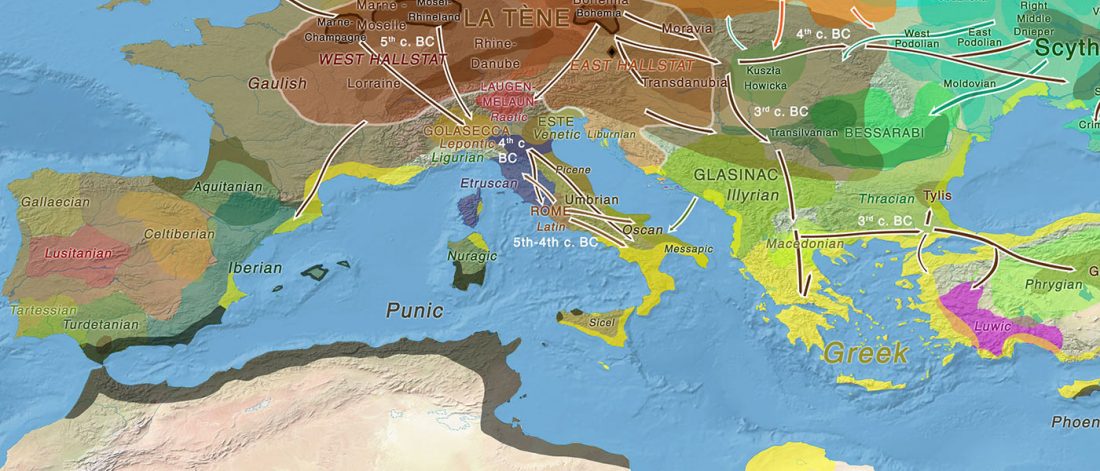
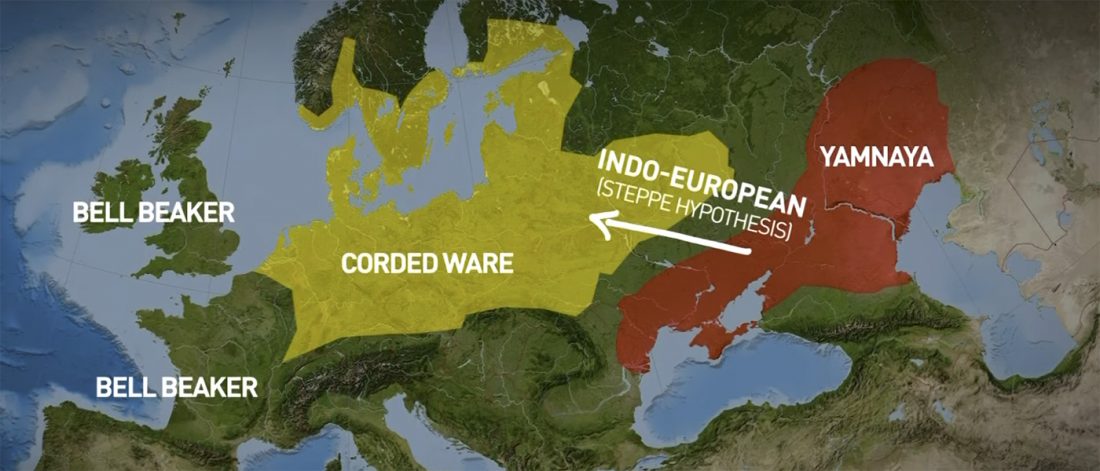
The genotypes from Human auditory ossicles as an alternative optimal source of ancient DNA, by Sirak et al. Genome Res. (2020), have been finally published by the Reich Lab, so we can get a sneak peek into what’s coming in future papers about the origins of R1a-rich Proto-Corded Ware and R1b-rich Italo-Venetic peoples.
NOTE. To avoid adding potential errors, I have merged the Reich Lab’s Curated Dataset (v. 42.4, March 1 2020) with these new samples before performing the qpAdm analyses. If you find something different with your files, you should probably check out this simple setting first. … Read the rest “Fully Steppe-like Proto-Corded Ware Late Trypillians”