The waves of disinformation are already here, putting the blame again on the European Union, as in the Financial Crisis of 2008. After years of negligent state policies promoted or tolerated by ruling political parties and social majorities of each country in the EU, which have led directly to yet another avoidable crisis. After years of state inactivity in the supranational political arena, hindering European social integration, and stripping EU institutions of any real power. The culprits are, again, not we, but they: evil and foreign hands pulling invisible strings from Brussels. The Age of Populism at its … Read the rest “Winds of change and our shared European past”
Tag: bell beaker
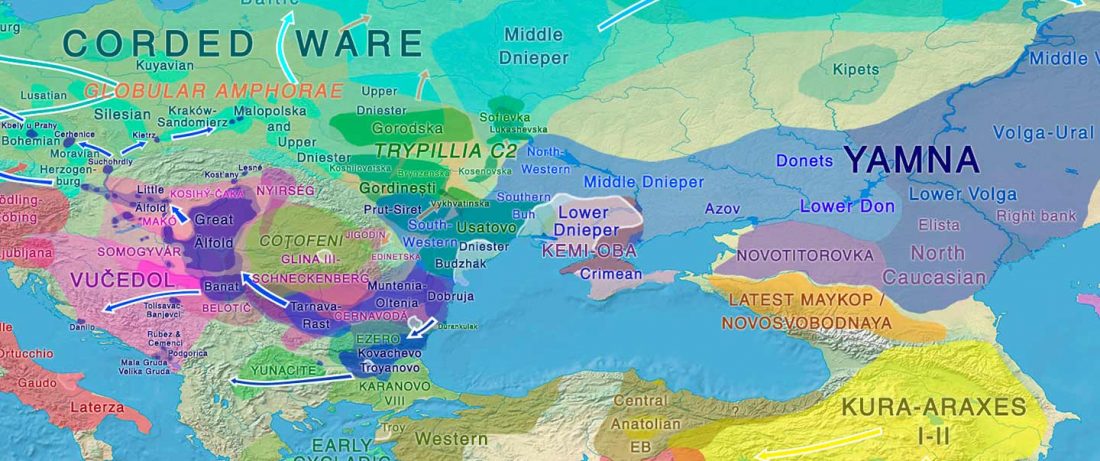
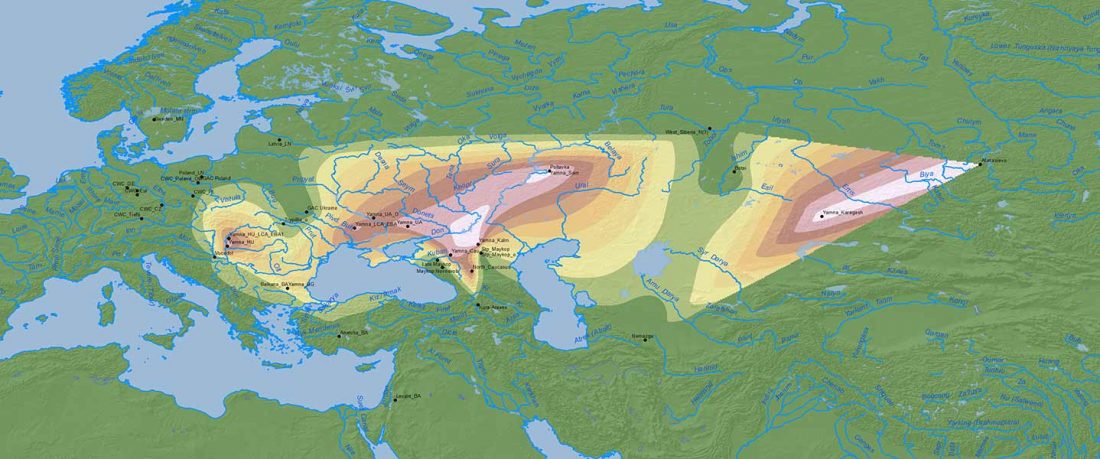
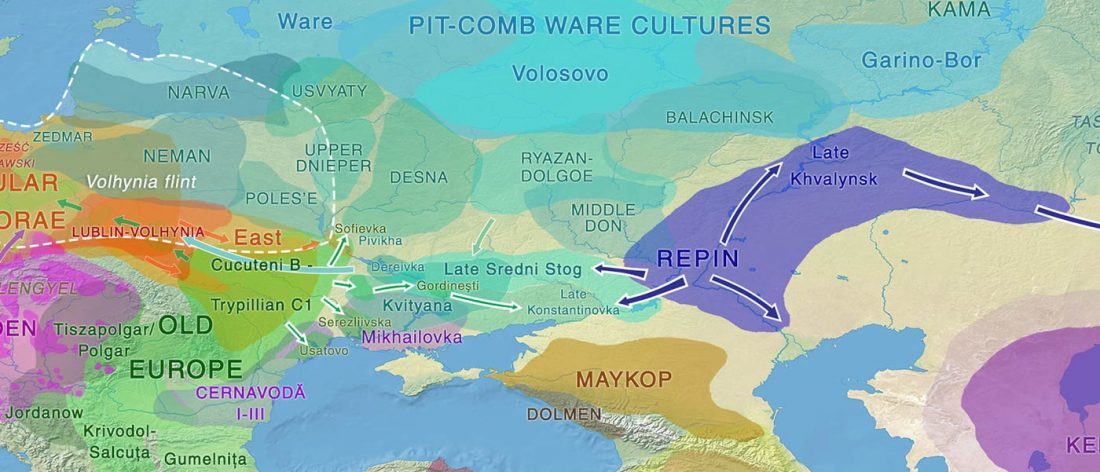
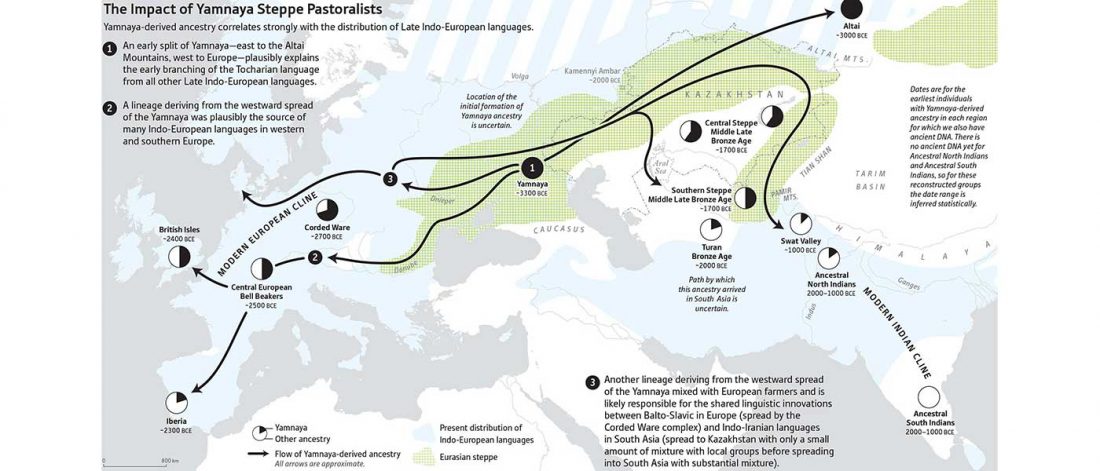
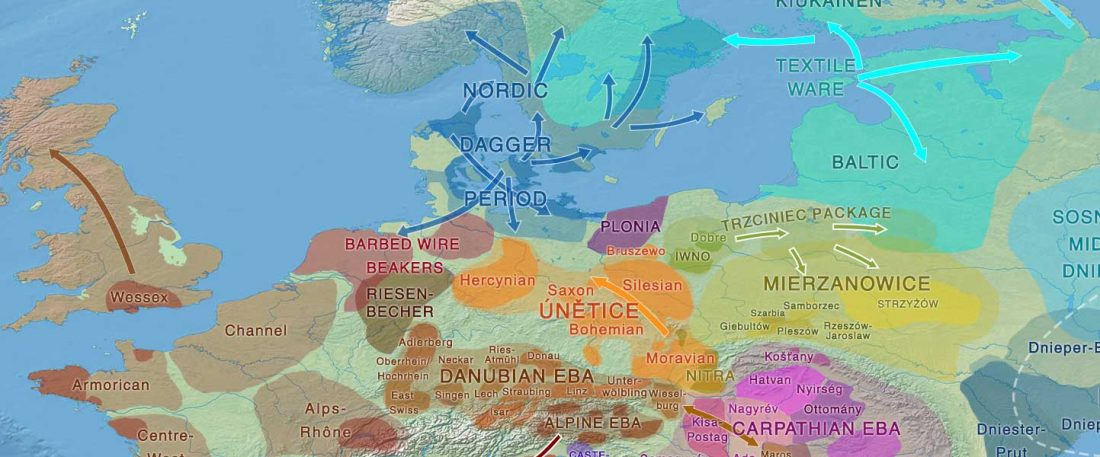
Fully Steppe-like Proto-Corded Ware Late Trypillians
The genotypes from Human auditory ossicles as an alternative optimal source of ancient DNA, by Sirak et al. Genome Res. (2020), have been finally published by the Reich Lab, so we can get a sneak peek into what’s coming in future papers about the origins of R1a-rich Proto-Corded Ware and R1b-rich Italo-Venetic peoples.
NOTE. To avoid adding potential errors, I have merged the Reich Lab’s Curated Dataset (v. 42.4, March 1 2020) with these new samples before performing the qpAdm analyses. If you find something different with your files, you should probably check out this simple setting first. … Read the rest “Fully Steppe-like Proto-Corded Ware Late Trypillians”
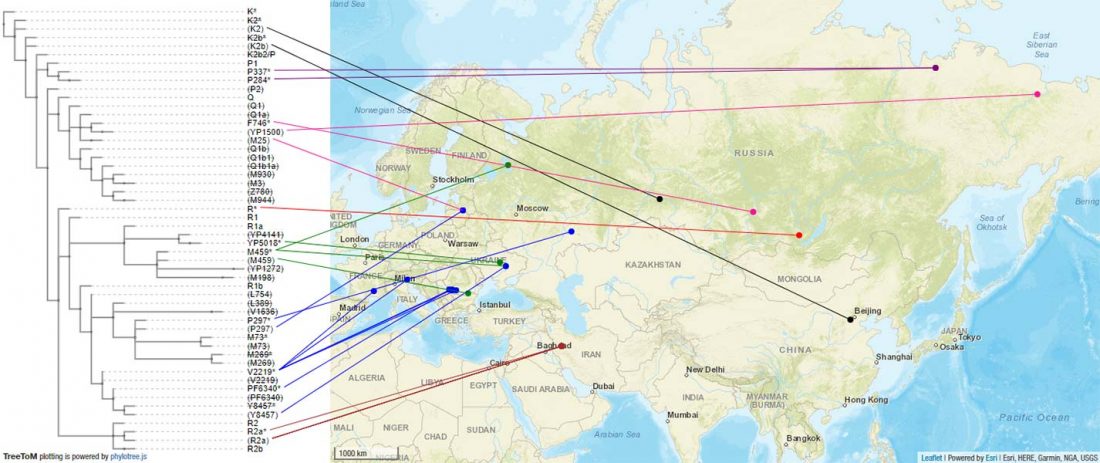
Ancient phylogeography: spread of haplogroups R1b, R1a and N
The previous post showed the potential use of TreeToM to visualize ancient DNA samples in maps together with their Y-DNA phylogenetic trees. I have written Newick trees for Y-chromosome haplogroups R1b-L388 (encompassing R-V1636 and R-P297, which in turn split into R-M73 and R-M269), R1a, and N.
I have reviewed some of the BAM files from my previous bulk analyses with YLeaf v.2, to add information that I had not previously included in the All Ancient DNA Dataset, and which might be relevant to the proper depiction of phylogenetic trees; in particular, positive and negative SNPs potentially distinguishing archaic… Read the rest “Ancient phylogeography: spread of haplogroups R1b, R1a and N”
R1b-L23-rich Bell Beaker-derived Italic peoples from the West vs. Etruscans from the East
New paper (behind paywall) Ancient Rome: A genetic crossroads of Europe and the Mediterranean, by Antonio et al. Science (2019).
The paper offers a lot of interesting data concerning the Roman Empire and more recent periods, but I will focus on Italic and Etruscan origins.
NOTE. I have updated prehistoric maps with Y-DNA and mtDNA data, and also the PCA of ancient Eurasian samples by period including the recently published samples, now with added sample names to find them easily by searching the PDFs.
Apennine homeland problem
The traditional question of Italic vs. Etruscan origins from a cultural-historical … Read the rest “R1b-L23-rich Bell Beaker-derived Italic peoples from the West vs. Etruscans from the East”
Corded Ware and Bell Beaker related groups defined by patrilocality and female exogamy
Two new interesting papers concerning Corded Ware and Bell Beaker peoples appeared last week, supporting yet again what is already well-known since 2015 about West Uralic and North-West Indo-European speakers and their expansion.
Below are relevant excerpts (emphasis mine) and comments.
#UPDATE (27 OCT 2019): I have updated Y-DNA and mtDNA maps of Corded Ware, Bell Beaker, EBA, MBA, and LBA migrations. I have also updated PCA plots, which now include the newly reported samples and those from the Tollense valley, and I have tried some qpAdm models (see below).
I. Corded Ware and
… Read the rest “Corded Ware and Bell Beaker related groups defined by patrilocality and female exogamy”Bell Beakers and Mycenaeans from Yamnaya; Corded Ware from the forest steppe
I have recently written about the spread of Pre-Yamnaya or Yamnaya ancestry and Corded Ware-related ancestry throughout Eurasia, using exclusively analyses published by professional geneticists, and filling in the gaps and contradictory data with the most reasonable interpretations. I did so consciously, to avoid any suspicion that I was interspersing my own data or cherry picking results.
Now I’m finished recapitulating the known public data, and the only way forward is the assessment of these populations using the available datasets and free tools.
Understanding the complexities of qpAdm is fairly difficult without a proper genetic and statistical background, which I … Read the rest “Bell Beakers and Mycenaeans from Yamnaya; Corded Ware from the forest steppe”
On the Ukraine Eneolithic outlier I6561 from Alexandria
Over the past week or so, since the publication of new Corded Ware samples in Narasimhan, Patterson et al. (2019) and after finding out that the R1a-M417 star-like phylogeny may have started ca. 3000 BC, I have been ruminating the relevance of contradictory data about the Ukraine_Eneolithic_o sample from Alexandria, its potential wrong radiocarbon date, and its implications for the Indo-European question.
How many other similar ‘controversial’ samples are there which we haven’t even considered? And what mechanisms are in place to control that the case of Hajji_Firuz_CA I2327 is not repeated?
Ukraine Eneolithic outlier I6561
It was not … Read the rest “On the Ukraine Eneolithic outlier I6561 from Alexandria”
Yamnaya replaced Europeans, but admixed heavily as they spread to Asia
Recent papers The formation of human populations in South and Central Asia, by Narasimhan, Patterson et al. Science (2019) and An Ancient Harappan Genome Lacks Ancestry from Steppe Pastoralists or Iranian Farmers, by Shinde et al. Cell (2019).
NOTE. For direct access to Narasimhan, Patterson et al. (2019), visit this link courtesy of the first author and the Reich Lab.
I am currently not on holidays anymore, and the information in the paper is huge, with many complex issues raised by the new samples and analyses rather than solved, so I will stick to the Indo-European question, … Read the rest “Yamnaya replaced Europeans, but admixed heavily as they spread to Asia”
Predictions about the genetic change from Single Grave to the Late Neolithic in Denmark
New open access paper Mapping human mobility during the third and second millennia BC in present-day Denmark by Frei et al. PLOS One (2019), from the Copenhagen group (including Allentoft, Sikora, and Kristiansen) of samples whose genomic profile will probably be published soon.
Interesting excerpts (emphasis mine):
… Read the rest “Predictions about the genetic change from Single Grave to the Late Neolithic in Denmark”We present results of the largest multidisciplinary human mobility investigation to date of skeletal remains from present-day Denmark encompassing the 3rd and 2nd millennia BC. Through a multi-analytical approach based on 88 individuals from 37 different archaeological localities in which we combine strontium isotope and radiocarbon analyses together with anthropological investigations, we explore