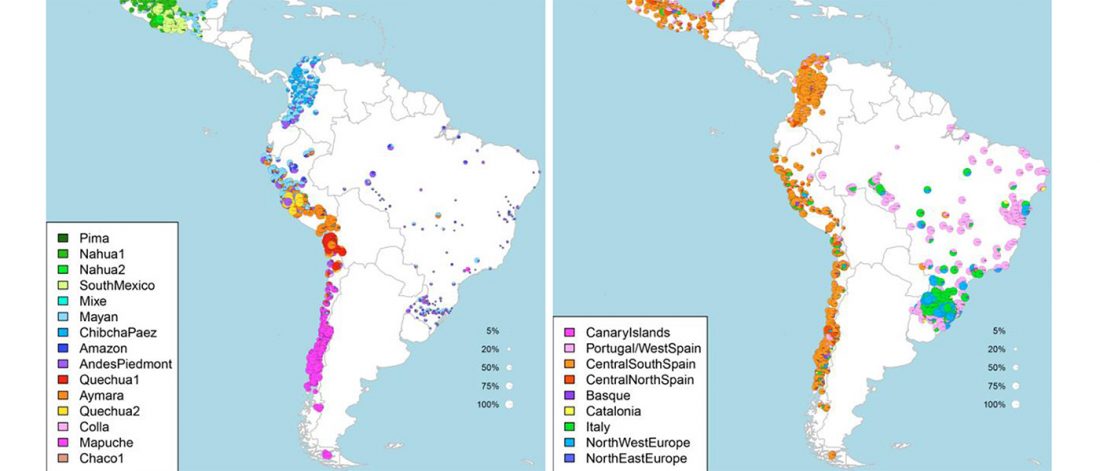
Latin Americans show widespread Mediterranean and North African ancestry
Recent preprint Latin Americans show wide-spread Converso ancestry and the imprint of local Native ancestry on physical appearance, by Chacon-Duque et al. bioRxiv (2018).
Abstract:
… Read the rest “Latin Americans show widespread Mediterranean and North African ancestry”Historical records and genetic analyses indicate that Latin Americans trace their ancestry mainly to the admixture of Native Americans, Europeans and Sub-Saharan Africans. Using novel haplotype-based methods here we infer the sub-populations involved in admixture for over 6,500 Latin Americans and evaluate the impact of sub-continental ancestry on the physical appearance of these individuals. We find that pre-Columbian Native genetic structure is mirrored in Latin Americans and that sources of non-Native ancestry, and admixture