Open access Global demographic history of human populations inferred from whole mitochondrial genomes, by Miller, Manica, and Amos, Royal Society Open Science (2018).
Relevant excerpts (emphasis mine):
Material
The Phase 3 sequence data from 20 populations, comprising five populations for each of the four main geographical regions of Europe, East Asia, South Asia and Africa, were downloaded from the 1000 Genomes Project website (www.1000genomes.org/data, [8]), including whole mitochondrial genome data for 1999 individuals. We decided not to analyse populations from the Americas due to the region’s complex history of admixture [13,14].
The European populations were as follows: Finnish sampled in Finland (FIN); European Caucasians resident in Utah, USA (CEU); British in England and Scotland (GBR); an Iberian population from Spain (IBS) and Toscani from Italy (TSI). Representing East Asia were the Han Chinese in Beijing (CHB); Southern Han Chinese (CHS); Dai Chinese from Xishuangbanna, China (CDX); Kinh population from Ho Chi Minh City, Vietnam (KHV) and Japanese from Tokyo (JPT). The South Asian populations were Punjabi Indians from Lahore, Pakistan (PJL); Gujarati Indians in Houston, USA (GIH) as well as Indian Telugu sampled in the UK (ITU); Bengali from Bangladesh (BEB) and Sri Lankan Tamil from the UK (STU). (…)
Method
We analysed our mtDNA data with the extended Bayesian skyline plot (EBSP) method, a Bayesian, non-parametric technique for inferring past population size fluctuations from genetic data. Building on the previous Bayesian skyline plot (BSP) approach, EBSP uses a piecewise-linear model and Markov chain Monte Carlo (MCMC) methods to reconstruct a populations’ demographic history [17] and is implemented in the software package BEAST v. 2.3.2 [11]. Alignments for each of the 20 populations were loaded separately into the Bayesian Evolutionary Analysis Utility tool (BEAUti v. 2.3.2) in NEXUS format.
Regional demographic histories
Europe:
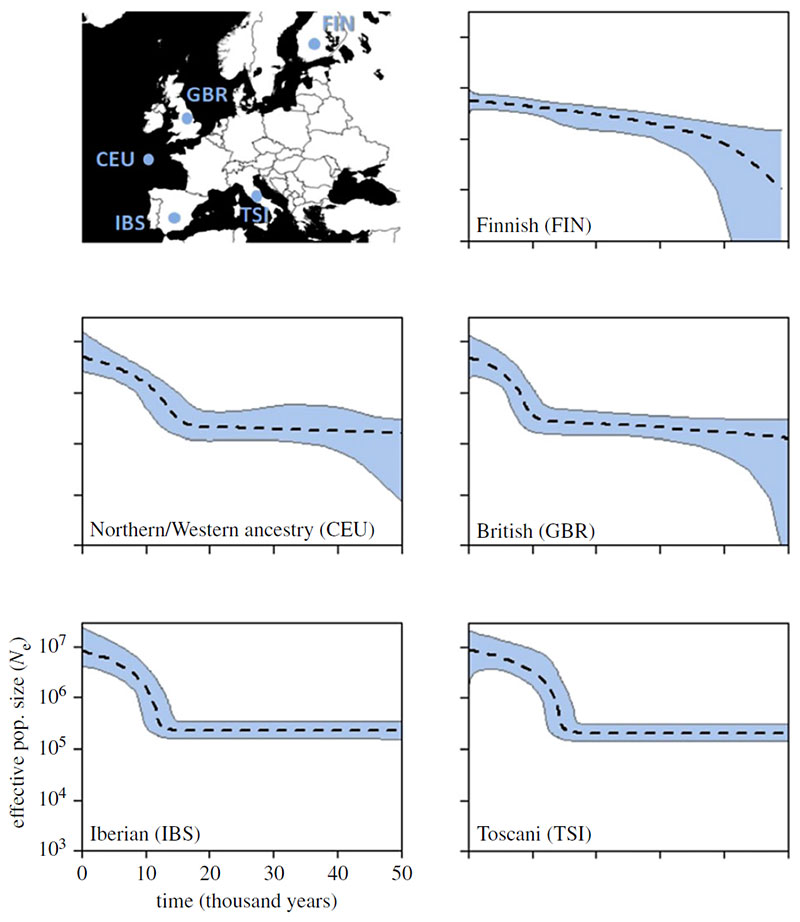
The five European profiles are presented in figure 2. The four southerly populations all show profiles with a stable size up to approximately 14 ka followed by a sudden, rapid increase that becomes progressively less steep towards the present. There is also a north-south trend, with confidence intervals becoming broader towards the north, particularly for the oldest time-points. The Finnish population profile appears rather different, but this is to be expected both because it is so far north and because previous studies have identified Finns as a strong genetic outlier in Europe [19–22].
South Asia:
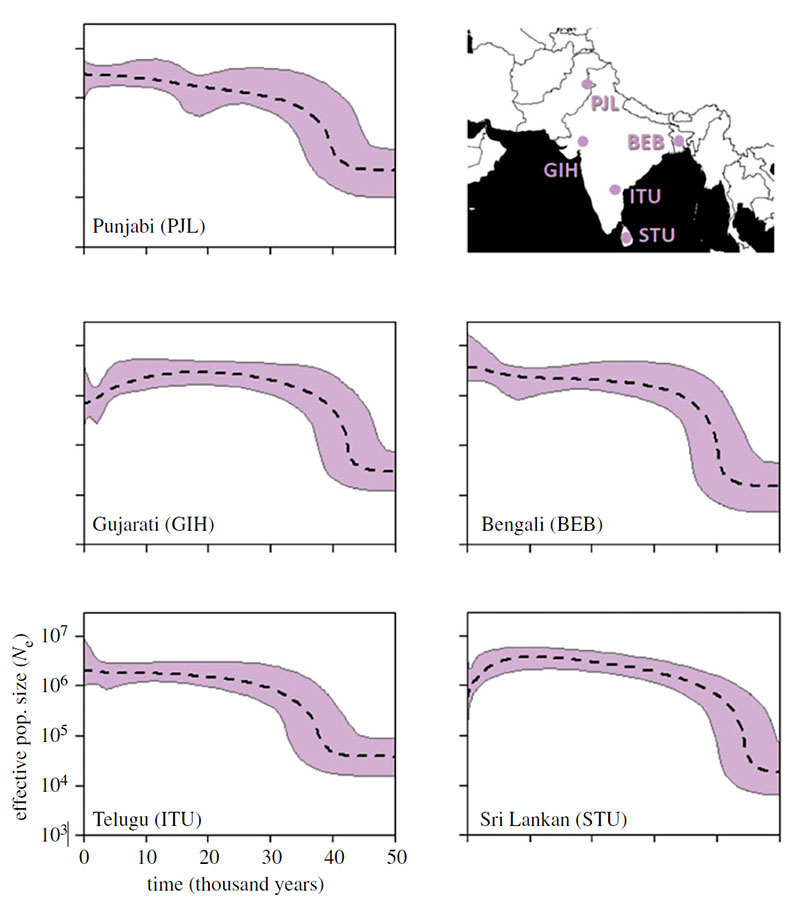
The five profiles for South Asia are shown in figure 3. All populations reveal a period of rapid growth approximately 45–40 ka which then slows. Near the present the two southerly populations, GIH and STU both show evidence of a decline. However, this may be due to these samples being drawn from populations no longer living on the subcontinent, with the downward trend capturing a bottleneck associated with moving to Europe/America, perhaps accentuated by the tendency for immigrant populations to group by region, religion and race [23].
Related
- Another hint at the role of Corded Ware peoples in spreading Uralic languages into north-eastern Europe, found in mtDNA analysis of the Finnish population
- Mitogenomes show continuity of Neolithic populations in Southern India
- North Asian mitogenomes hint at the arrival of pastoralists from West to East ca. 2800-1000 BC
- Demographic history and genetic adaptation in the Himalayan region
- Mitogenomes from Thailand offer insights into maternal genetic history of mainland South-East Asia
- Hungarian mitogenomes similar to East and West Slavs, but genetic substratum predates their historic contacts