New paper (behind paywall) Neolithic phylogenetic continuity inferred from complete mitochondrial DNA sequences in a tribal population of Southern India, by Sylvester et al. Genetica (2018).
This paper used a complete mtDNA genome study of 113 unrelated individuals from the Melakudiya tribal population, a Dravidian speaking tribe from the Kodagu district of Karnataka, Southern India.
Some interesting excerpts (emphasis mine):
Autosomal genetic evidence indicates that most of the ethnolinguistic groups in India have descended from a mixture of two divergent ancestral populations: Ancestral North Indians (ANI) related to People of West Eurasia, the Caucasus, Central Asia and the Middle East, and Ancestral South Indians (ASI) distantly related to indigenous Andaman Islanders (Reich et al. 2009). It is presumed that proto-Dravidian language, most likely originated in Elam province of South Western Iran, and later spread eastwards with the movement of people to the Indus Valley and later the subcontinent India (McAlpin et al. 1975; Cavalli-Sforza et al. 1988; Renfrew 1996; Derenko et al. 2013). West Eurasian haplogroups are found across India and harbor many deep-branching lineages of Indian mtDNA pool, and most of the mtDNA lineages of Western Eurasian ancestry must have a recent entry date less than 10 Kya (Kivisild et al. 1999a). The frequency of these lineages is specifically found among the higher caste groups of India (Bamshad et al. 1998, 2001; Basu et al. 2003) and many caste groups are direct descendants of Indo-Aryan immigrants (Cordaux et al. 2004). These waves of various invasions and subsequent migrations resulted in major demographic expansions in the region, which added new languages and cultures to the already colonized populations of India. Although previous genetic studies of the maternal gene pools of Indians had revealed a genetic connection between Iranian populations and the Arabian Peninsula, likely the result of both ancient and recent gene flow (Metspalu et al. 2004; Terreros et al. 2011).
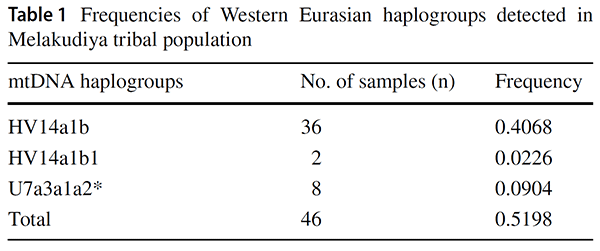
Haplogroup HV14
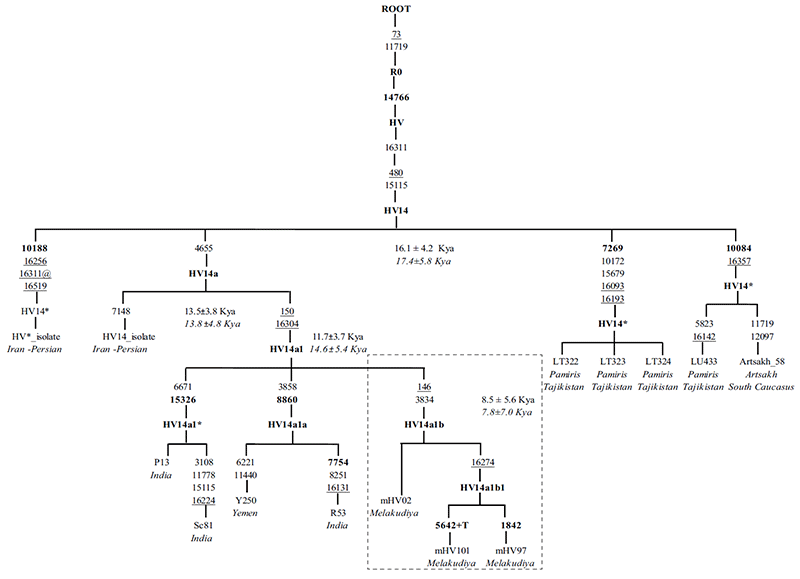
mtDNA haplogroup HV14 has prominence in North/Western Europe, West Eurasia, Iran, and South Caucasus to Central Asia (Malyarchuk et al. 2008; Schonberg et al. 2011; Derenko et al. 2013; De Fanti et al. 2015). Although Palanichamy identified haplogroup HV14a1 in three Indian samples (Palanichamy et al. 2015), it is restricted to limited unknown distribution. In the present study, by the addition of considerable sequences from the Melakudiya population, a unique novel subclade designated as HV14a1b was found with a high frequency (43%) allowed us to reveal the earliest diverging sequences in the HV14 tree prior to the emergence of HV14a1b in Melakudiya. (…) The coalescence age for haplogroup HV14 in this study is dated ~ 16.1 ± 4.2 kya and the founder age of haplogroup HV14 in Melakudiya tribe, which is represented by a novel clade HV14a1b is ~ 8.5 ± 5.6 kya
Haplogroup U7a3a1a2
The coalescence age of haplogroup U7a3a1a2 dates to ~ 13.3 ± 4.0 kya. (…)
Although, haplogroup U7 has its origin from the Near East and is widespread from Europe to India, the phylogeny of Melakudiya tribe with subclade U7a3a1a2 clusters with populations of India (caste and tribe) and neighboring populations (Irwin et al. 2010; Ranaweera et al. 2014; Sahakyan et al. 2017), hint about the in-situ origin of the subclade in India from Indo-Aryan immigrants.
I am not a native English speaker, but this paper looks like it needs a revision by one.
Also – without comparison with ancient DNA – it is not enough to show coalescence age to prove an origin of haplogroup expansion in the Neolithic instead of later bottlenecks. However, since we are talking about mtDNA, it is likely that their analysis is mostly right.
Finally, one thing is to prove that the origin of the Indus Valley Civilization lies (in part) in peoples from the Iranian plateau, and to show with ASI ancestry that they are probably the origin of Proto-Dravidian expansion, and another completely different thing is to prove an Elamo-Dravidian connection.
Since that group is not really accepted in linguistics, it is like talking about proving – through that Iran Neolithic ancestry – a Sumero-Dravidian, or a Hurro-Dravidian connection…
Related
- Mitogenomes show ancient human migrations to and through North-East India not of males exclusively
- The Indus Valley Civilisation in genetics – the Harappan Rakhigarhi project
- The Aryan migration debate, the Out of India models, and the modern “indigenous Indo-Aryan” sectarianism
- Eurasian steppe dominated by Iranian peoples, Indo-Iranian expanded from East Yamna
- Early Indo-Iranian formed mainly by R1b-Z2103 and R1a-Z93, Corded Ware out of Late PIE-speaking migrations
- Y-DNA haplogroup R1b-Z2103 in Proto-Indo-Iranians?
- Indo-European and Central Asian admixture in Indian population, dependent on ethnolinguistic and geodemographic divisions
