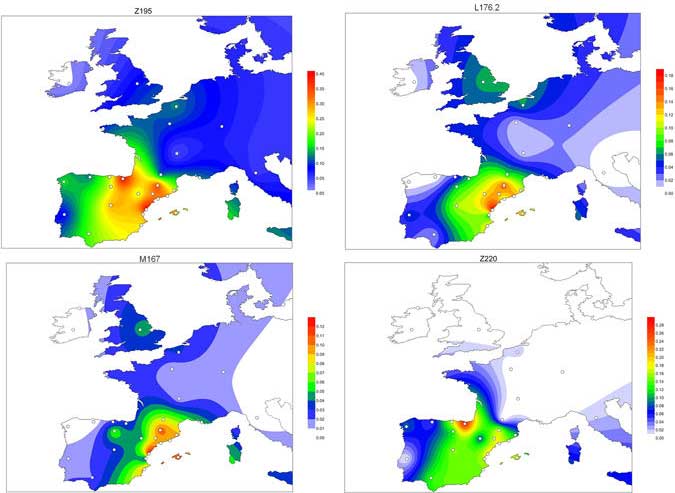
As you can see from my interest in the recently published Olalde et al. (2019) Iberia paper, once you accept that East Bell Beakers expanded North-West Indo-European, the most important question becomes how did its known dialects spread to their known historic areas.
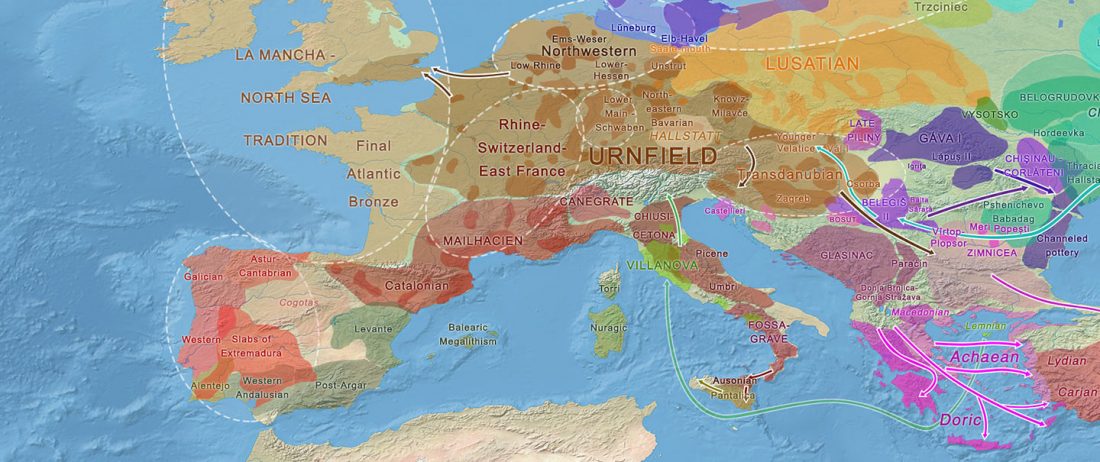
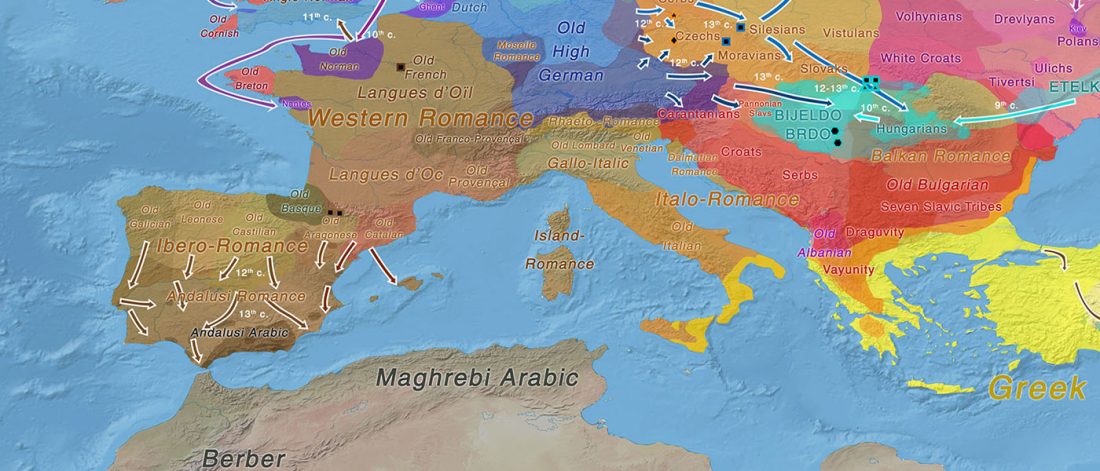
We already had a good idea about the expansion of Celts, based on proto-historical accounts, fragmentary languages, and linguistic guesstimates, but the connection of Celtic with either Urnfield or slightly later Hallstatt/La Tène was always blurred, due to the lack of precise data on population movements.
The latest paper on Iberia is interesting for many … Read the rest “Haplogroup R1b-M167/SRY2627 linked to Celts expanding with the Urnfield culture”