The following are some recent developments and updates:
I. Ancient DNA Dataset version 2
I.1. Accurate mtDNA haplogroups
I was meaning to update the mtDNA part of the Ancient DNA Dataset, and finally found some time to review FTDNA and YFull nomenclature (including hyperlinks), as well as those SNP calls from published samples found in YFull’s MTree. So, if you are interested in studies of mtDNA phylogeography, I think the data is now accurate and much more useful.
Given the number of columns and the size of the files, I have decided to post shorter standard versions, by cutting the columns used mostly for GIS, leaving them only in the version labelled full. See the Google Drive folder.
I.2. Lactase persistence evolution
I have also added a new column LP with the recent full reports on lactase persistence from Segurel et al. PLOS Biology (2020), which continues Iain Mathieson’s work on the subject. As expected, except for the Alexandria outlier (a likely misdated sample due to a reservoir effect), the allele seems to spread only after the westward expansion of the Yamnaya.
See a spreadsheet with samples showing a 0/1 result.
Some Early Bronze Age populations show a noticeable LP prevalence, including Bell Beakers and derived groups in Central Europe, and Steppe-related populations in Asia, which suggests that this trait might have been linked to admixture events of the Yamnaya with an EEF-related population of the north-west Pontic area or the Balkans.
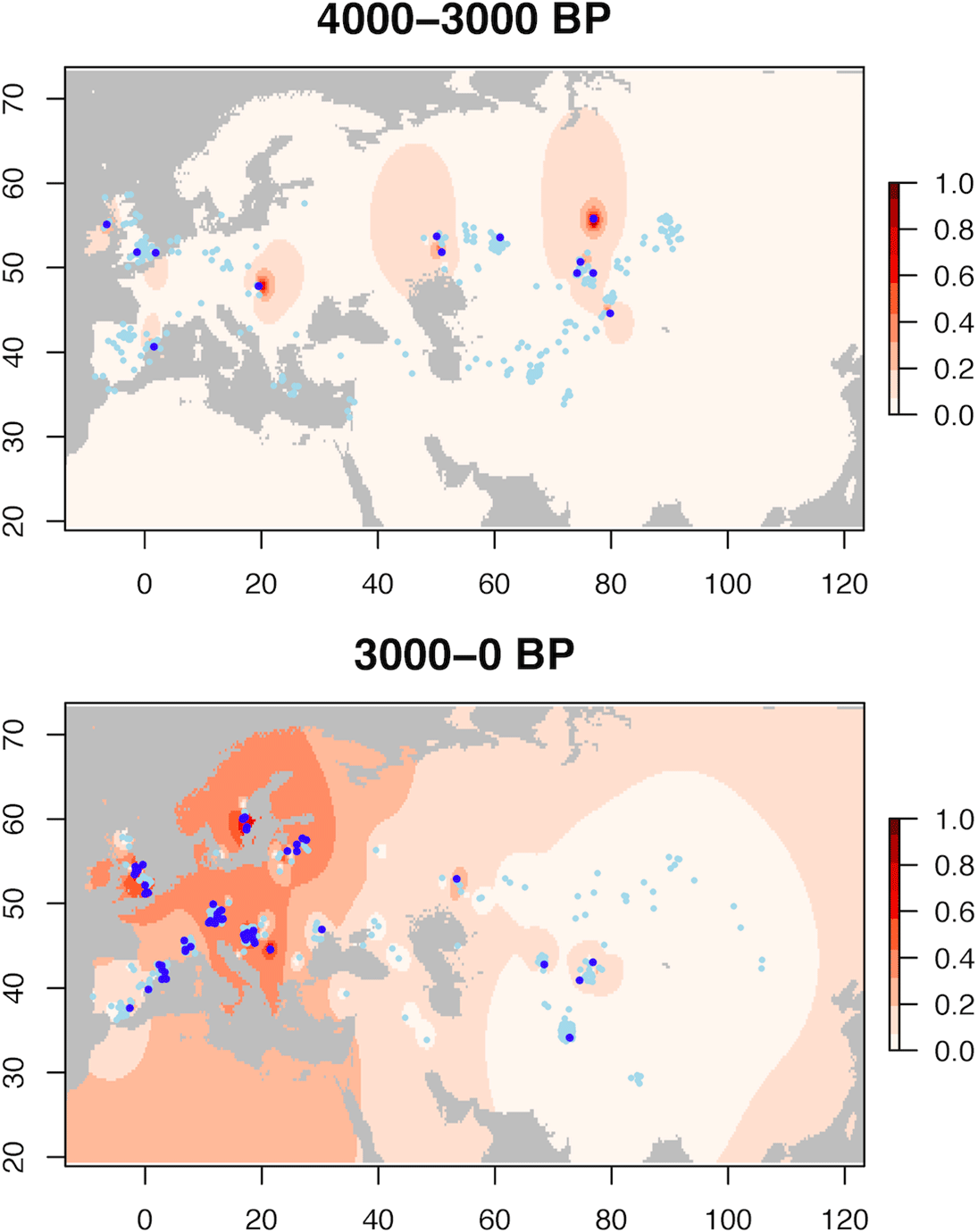
Here is a video with a timeline and geographical progression based on the published data:
Timing and geographical progression of lactase persistent vs. lactase nonpersistent individuals from the Mesolithic to the Middle Ages: black circle = 0; red star = 1; others = cross. Date on bottom right corner. Data from https://t.co/dQia3G5s2q pic.twitter.com/3WzRM9nhrx
— Carlos Quiles (@cquilesc) June 15, 2020
Here is the corresponding OpenLayers map (might be slow to load):
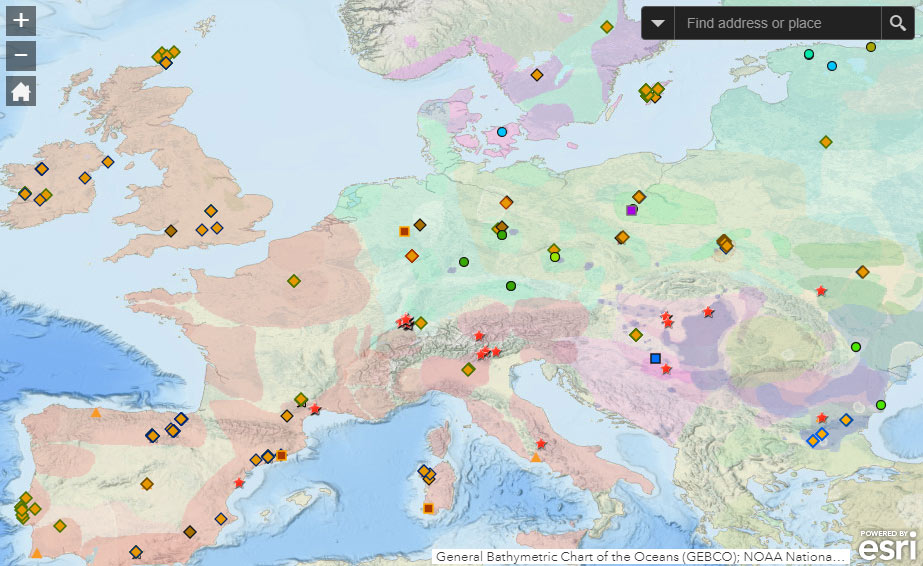
![]() Geographical distribution of lactase persistence in ancient samples.
Geographical distribution of lactase persistence in ancient samples.
I.3. Y-DNA symbols
I have used the opportunity to review some Y-DNA symbols in the Web App, in particular subclade colors and borders (such as I1 and I2 subclades, or J2b-L283) so that their evolution can be more easily followed visually:
II. admixr: R for ADMIXTOOLS
If you followed the instructions for ancestry tools (2017) from this site, you probably noticed that I cut it short in the ADMIXTOOLS section due to some failures with a virtual Linux setting and the computer I used back then.
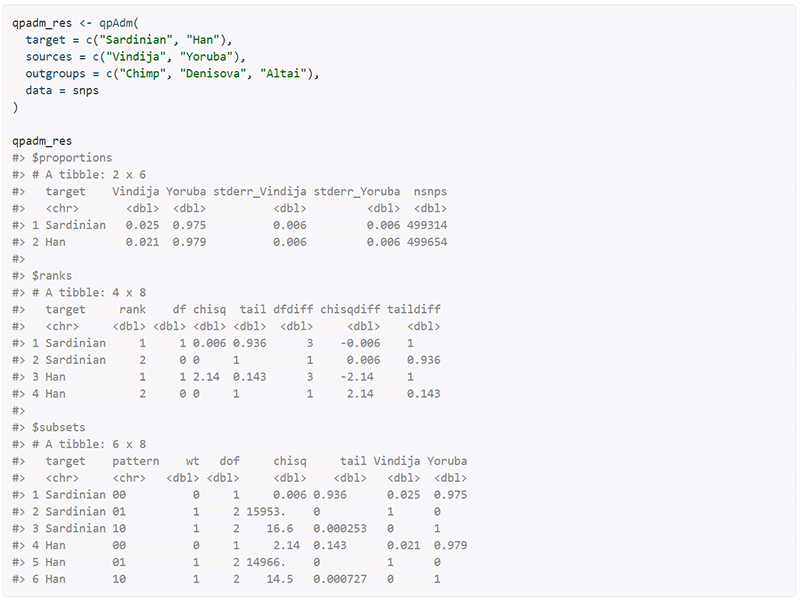
I made an interesting discovery in early 2019, which helped me get a bunch of qpAdm analyses done without disrupting my usual work with Windows: By using the R package admixr – which accesses ADMIXTOOLS and parses its output – I could leave my failing, noisy old PC with Linux at nights and get lots of processed results the next day.
Some of you might be thinking that a complex batch script would offer the same result as admixr. While this is true, especially for those used to manually work with the necessary files in Linux all the time, there is no comparison with admixr for the rest of us when it comes to automating the pipeline with dozens of models. Even better, admixr lets you group samples, perform merging and filtering, and indeed analyses such as F3 and F4 statistics, all from R. In any case, I’d recommend to give it first a try and then decide for yourself.
I had been waiting for the author to release an announced update, which would incorporate the latest recommendations of the Reich Lab on qpAdm best uses, and apparently that update is here as version 0.8.0:
I can't believe it's been a year since the last admixr release. It's not dead! A new version is out. As requested by a user on Twitter on Friday (and many others over time), you can now examine the full log behind any admixr function from R. #rstats https://t.co/KtdkMStwJZ
— Martin Petr (@fleventy5) June 14, 2020
On top of its corrections of minor errors, the new version has added parsing and saving of the log file, which until now had to be done by indirectly accessing the temp folder in R.
#EDIT (16 June 2020): Martin Petr has released new versions, partly due to comments received, including incorporating the default qpAdm parameters as recommended by the Reich Lab. If you think you might be interested in testing it, right now would be a great moment to contact him suggesting improvements, before the author moves on to other projects:
So, uh, for anyone still following: `devtools::install_github("bodkan/admixr", ref = "v0.8.7")` will give you expanded qpAdm reports, many new bugfixes (issues list nearly empty now!). I expect 0.9.0 will have basic automated qpAdm model fitting (see preprint by Harney et al.).
— Martin Petr (@fleventy5) June 15, 2020
As you can see from my posts when I publish qpAdm results, one of the great benefits I find in using admixr is to be able to copy&paste the simplified parsed results as tables in a spreadsheet. Similarly, when you want to test dozens of different models with only minor differences among them, you can organize all source and target populations by using any spreadsheet or text editor and copy&pasting as many combinations as you can imagine, leaving the tests going on for hours or days.
Rstudio online
During the confinement, I haven’t been able to use my study that much, because the missus confiscated my workstation to teach online at any given hour. I had to get creative if I wanted to access the computer, so I installed Rstudio to be able to use R in that old PC through the intranet from anywhere in the house, whether with a smartphone, tablet or laptop.
In case you are interested, it shouldn’t be difficult to install Rstudio on some online server (say, AWS at Amazon) and perform analyses fairly quickly. In fact, you could offer that kind of setting for others by using a Shiny app with a simple interface, such as by including a map to select samples from. I don’t know if that kind of project would be profitable, but offering access to professional tools would at least be a huge improvement over the countless amateur calculator- or coordinate-based games available out there…
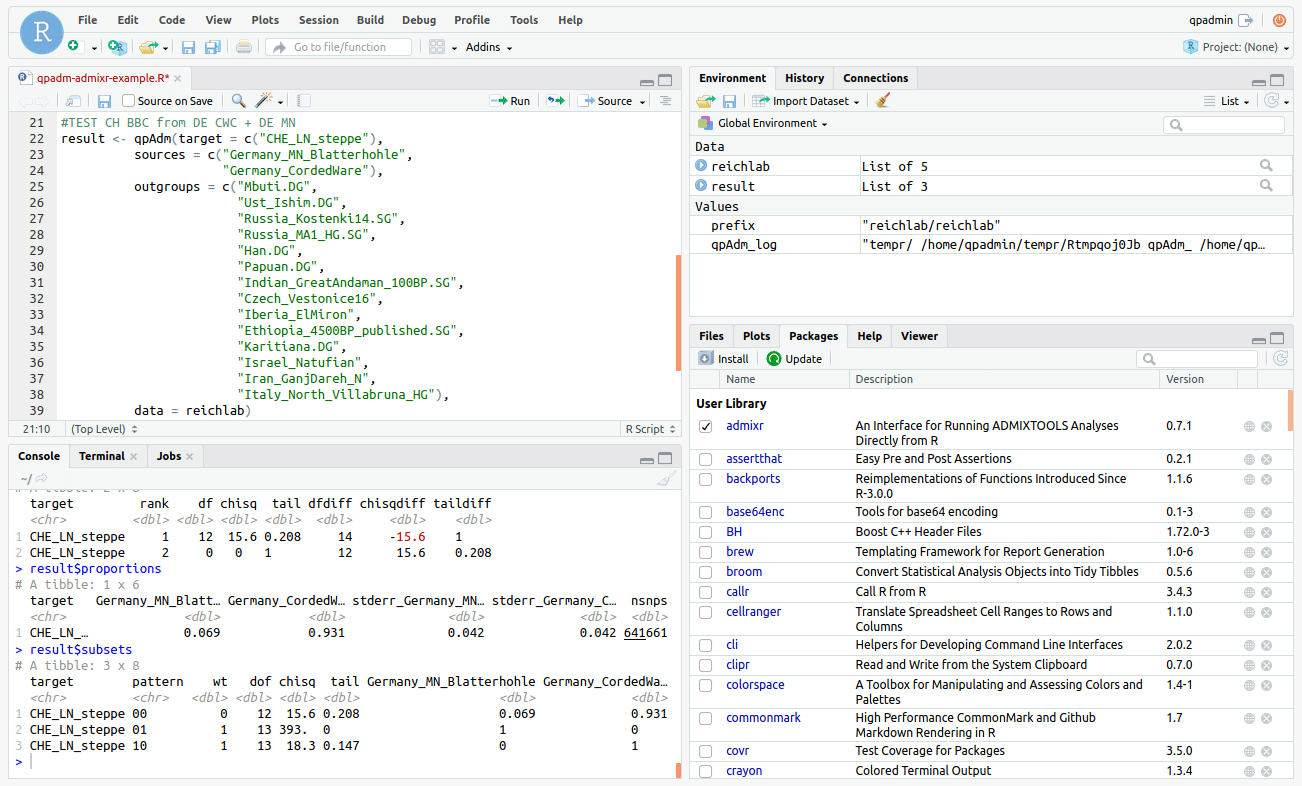
Now, since I don’t have much time lately, and I don’t really use ADMIXTOOLS except occasionally to test something very specific, I think it is only fair to offer any of you out there with a genuine interest in it – and without access to a proper setting – the possibility of working with Rstudio online (with an interface like the above) and without a cost.
NOTE. The merged genotype file is what I have at each given moment. For example, today’s one incorporates the Reich Lab’s curated dataset and most recently published samples.
Just write me an email at [email protected] and I will give you access (it only works with one Internet user at a time), provided that you are:
- Reasonable: preferably a long-time reader, with some previous comments and/or email, or else with some online history I can check out. That’s necessary because you would be accessing one of my computers.
- Knowledgeable: previous experience with ADMIXTOOLS would be great to avoid spending too much time asking me questions, time that could be dedicated to help improve the setting instead, now that I have direct access to the PC again and thus little interest in maintaining this.
The instructions for the use of admixr are fairly simple as they are.
I don’t care about your specific ideas or planned uses, whether you want to find out that Bell Beakers came from Iberia or Denmark, or Corded Ware from Yamnaya or Starcevo, or Yamnaya from Sredni Stog or Maykop, or Anatolia Neolithic from India or Central Asia…I really don’t care, because what is needed to solve questions is not some groundbreaking discovery in amateur ancestry magic, but data.
Using ADMIXTOOLS as an amateur should be mostly a way to try and reproduce results from published papers, and to better understand the advances and limitations of new reports.
#UPDATE (19 June 2019): Other researchers have different ways to automate qpAdm runs. Pontus Skoglund just released qpAdm_wrapper on GitHub – a Python script to cycle through sources:
Ancestry modellers: I have made my qpAdm_wrapper tool available https://t.co/BxXmeBbPu2, which we have been using in the lab. It allows Nick Patterson's qpAdm to be run on single command lines, and automatic "cycling" through all possible admixture sources as we did in Cell 2017.
— Pontus Skoglund (@pontus_skoglund) June 17, 2020