New paper (behind paywall) Ancient goat genomes reveal mosaic domestication in the Fertile Crescent, by Daly et al. Science (2018) 361(6397):85-88.
Interesting excerpts (emphasis mine):
Thus, our data favor a process of Near Eastern animal domestication that is dispersed in space and time, rather than radiating from a central core (3, 11). This resonates with archaeozoological evidence for disparate early management strategies from early Anatolian, Iranian, and Levantine Neolithic sites (12, 13). Interestingly, our finding of divergent goat genomes within the Neolithic echoes genetic investigation of early farmers. Northwestern Anatolian and Iranian human Neolithic genomes are also divergent (14–16), which suggests the sharing of techniques rather than large-scale migrations of populations across Southwest Asia in the period of early domestication. Several crop plants also show evidence of parallel domestication processes in the region (17).
PCA affinity (Fig. 2), supported by qpGraph and outgroup f3 analyses, suggests that modern European goats derive from a source close to the western Neolithic; Far Eastern goats derive from early eastern Neolithic domesticates; and African goats have a contribution from the Levant, but in this case with considerable admixture from the other sources (figs. S11, S16, and S17 and tables S26 and 27). The latter may be in part a result of admixture that is discernible in the same analyses extended to ancient genomes within the Fertile Crescent after the Neolithic (figs. S18 and S19 and tables S20, S27, and S31) when the spread of metallurgy and other developments likely resulted in an expansion of inter-regional trade networks and livestock movement.
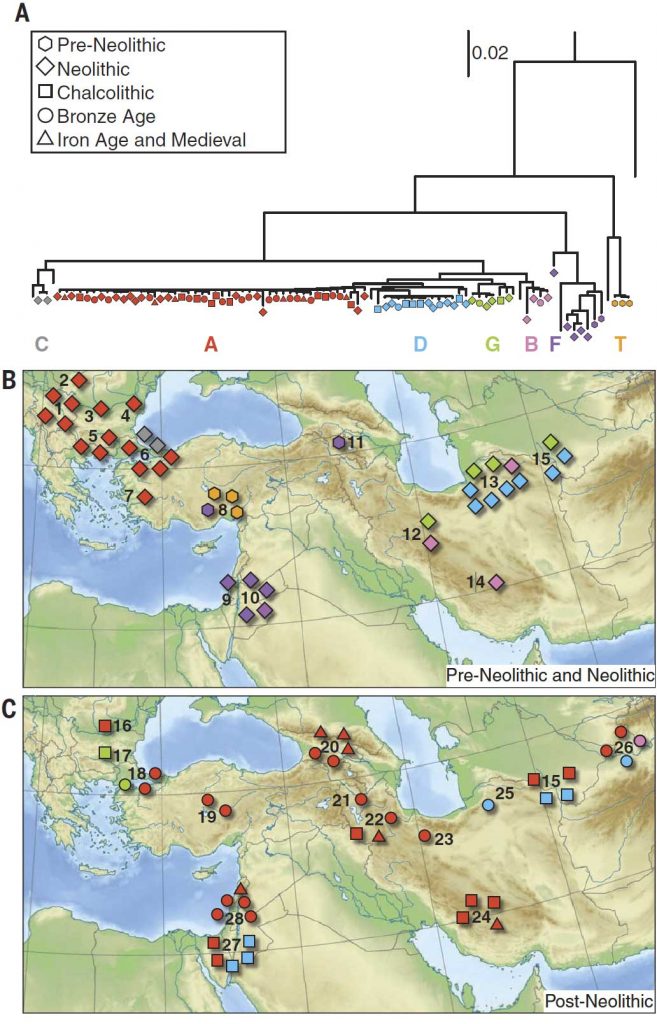
Our results imply a domestication process carried out by humans in dispersed, divergent, but communicating communities across the Fertile Crescent who selected animals in early millennia, including for pigmentation, the most visible of domestic traits.
Related
- Improving environmental conditions favoured higher local population density, which favoured domestication
- Domestication spread probably via the North Pontic steppe to Khvalynsk… but not horse riding
- About Scepters, Horses, and War: on Khvalynsk migrants in the Caucasus and the Danube
- Differing modes of animal exploitation in North Pontic Eneolithic and Bronze Age Societies
- North Pontic steppe Eneolithic cultures, and an alternative Indo-Slavonic model
- The Lower Danube during the Eneolithic, and the potential Proto-Anatolian community
- Eneolithic Ukraine cultures of the North Pontic steppe and southern steppe-forest, on the Left Bank of the Dnieper
- Recent archaeological finds near Indo-European and Uralic homelands
- Proto-Indo-European homeland south of the Caucasus?
- On the potential origin of Caucasus hunter-gatherer ancestry in Eneolithic steppe cultures
- Consequences of O&M 2018 (II): The unsolved nature of Suvorovo-Novodanilovka chiefs, and the route of Proto-Anatolian expansion