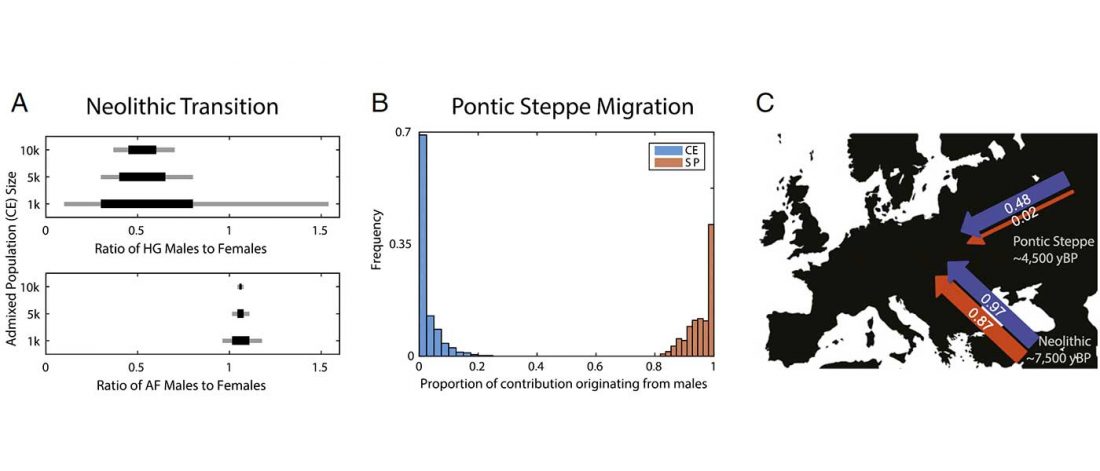
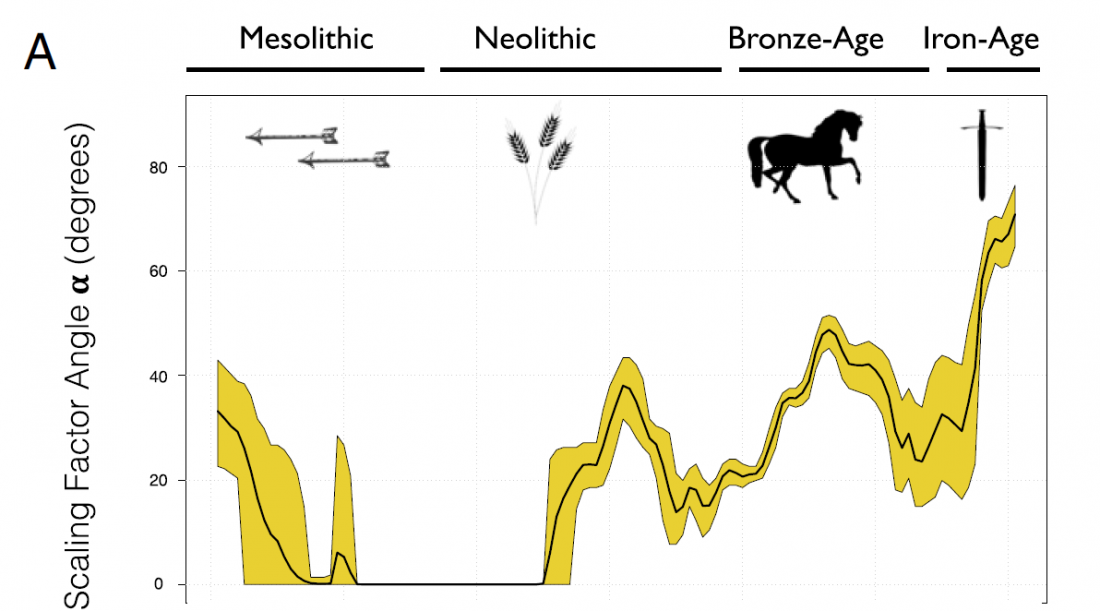
Two interesting papers questioning previous methods have been published.
Open access Mitochondrial DNA is unsuitable to test for isolation by distance, by Teske et al. Scientific Reports (2018) 8:8448.
Abstract (emphasis mine):
… Read the rest “Mitochondrial DNA unsuitable to test for IBD, and undersampling genomes show biased time and rate estimates”Tests for isolation by distance (IBD) are the most commonly used method of assessing spatial genetic structure. Many studies have exclusively used mitochondrial DNA (mtDNA) sequences to test for IBD, but this marker is often in conflict with multilocus markers. Here, we report a review of the literature on IBD, with the aims of determining (a) whether significant IBD is primarily a result of lumping spatially discrete