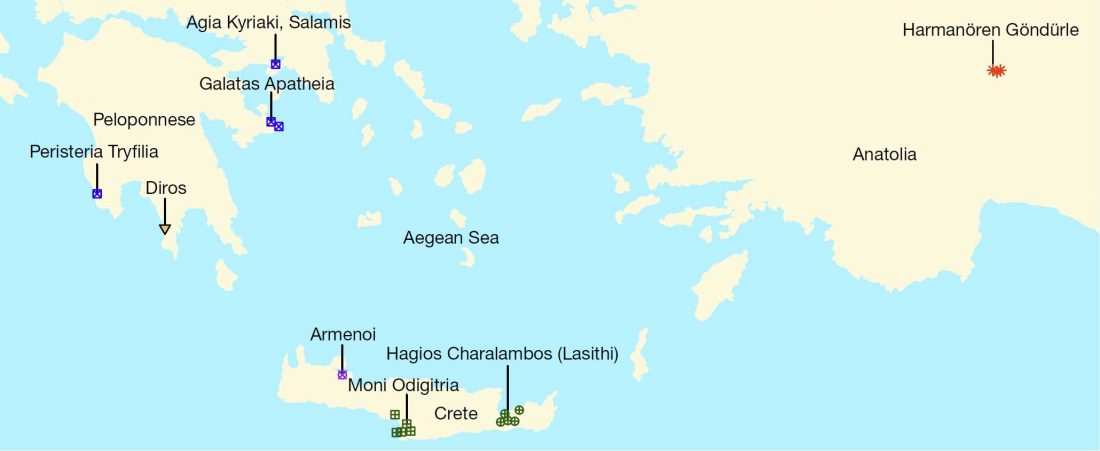
Genetic origins of Minoans and Mycenaeans and their continuity into modern Greeks
A new article has appeared in Nature, Genetic origins of the Minoans and Mycenaeans, by Lazaridis et al. (2017), referenced by Science.
Abstract:
… Read the rest “Genetic origins of Minoans and Mycenaeans and their continuity into modern Greeks”The origins of the Bronze Age Minoan and Mycenaean cultures have puzzled archaeologists for more than a century. We have assembled genome-wide data from 19 ancient individuals, including Minoans from Crete, Mycenaeans from mainland Greece, and their eastern neighbours from southwestern Anatolia. Here we show that Minoans and Mycenaeans were genetically similar, having at least three-quarters of their ancestry from the first Neolithic farmers of western Anatolia and the Aegean, and most of the remainder