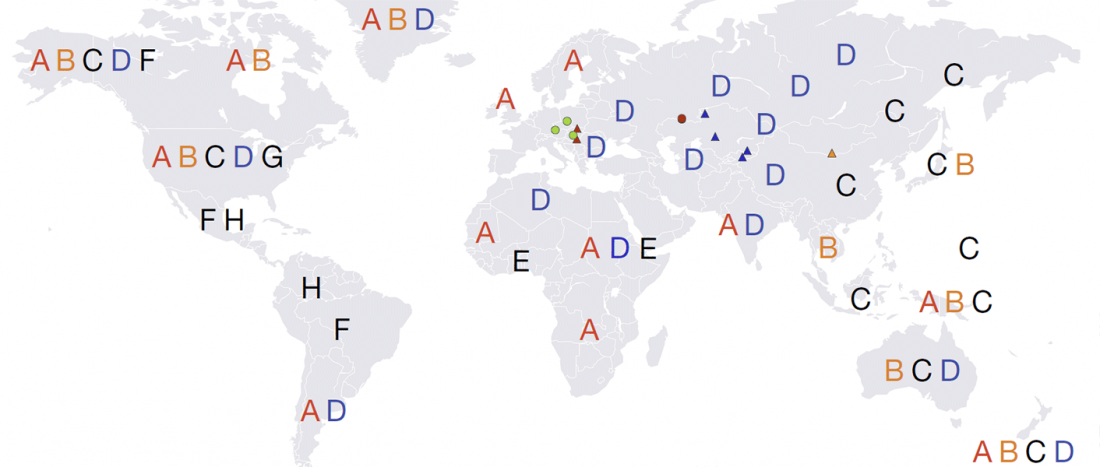
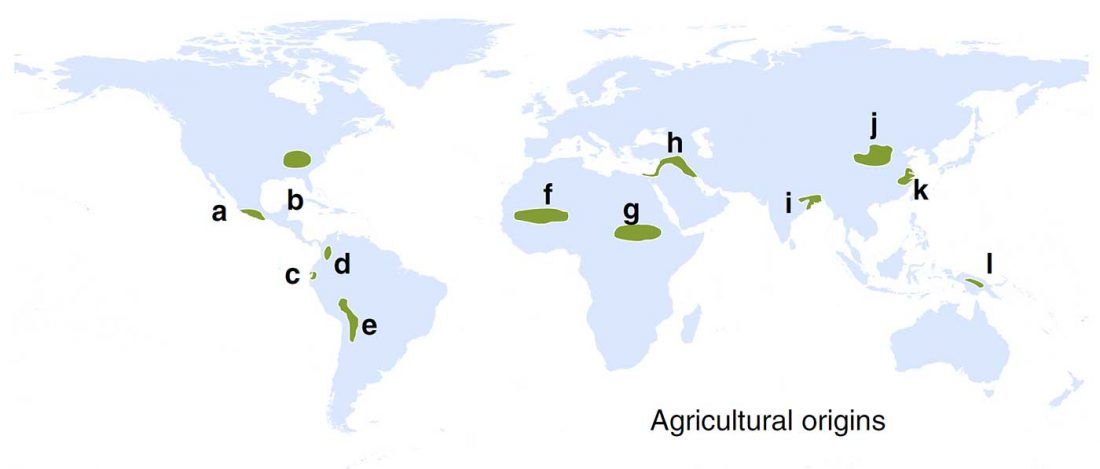
Ancient hepatitis B viruses from the Bronze Age to the Medieval period, by Mühlemann et al., Science (2018) 557:418–423.
NOTE. You can read the PDF at Dalia Pokutta’s Academia.edu account.
Abstract (emphasis):
… Read the rest “Shared ancestry of ancient Eurasian hepatitis B virus diversity linked to Bronze Age steppe”Hepatitis B virus (HBV) is a major cause of human hepatitis. There is considerable uncertainty about the timescale of its evolution and its association with humans. Here we present 12 full or partial ancient HBV genomes that are between approximately 0.8 and 4.5 thousand years old. The ancient sequences group either within or in a sister relationship with extant human or other ape HBV clades. Generally,