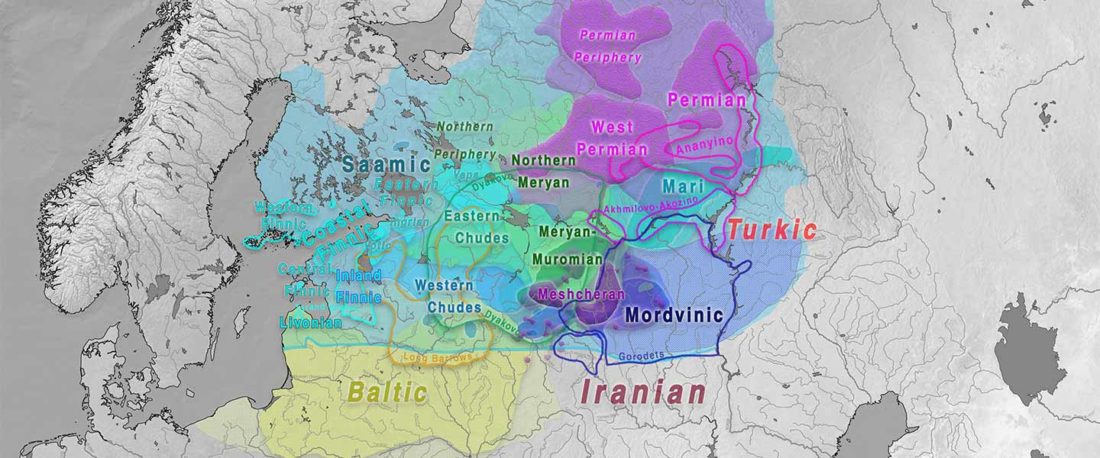
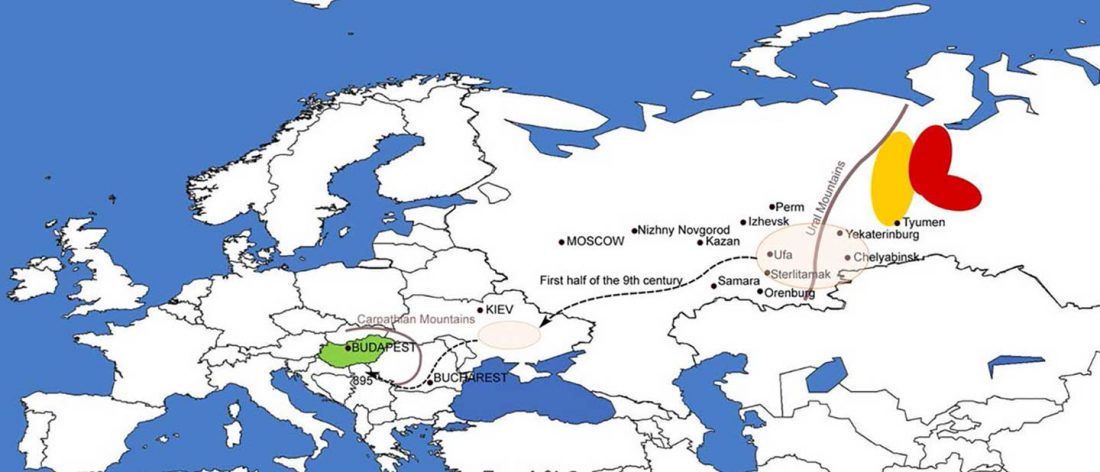
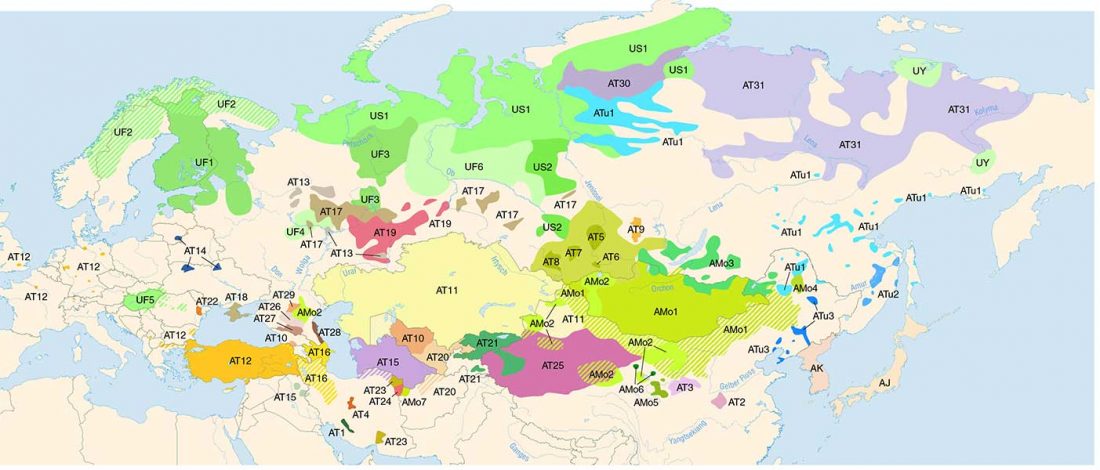
On the ethnolinguistic origin of the Meshchera, Pauli Rahkonen had an interesting proposal that might eventually be tested with Bronze Age and Iron Age DNA samples from North-East Europe: The Linguistic Background of the Ancient Meshchera Tribe and Principal Areas of Settlement, FUF (2009) 60:160-200.
NOTE. The paper is included in his PhD Dissertation, South-eastern contact area of Finnic languages in the light of onomastics (2013).
Interesting excerpts (emphasis mine, minor changes for clarity)
… Read the rest “European hydrotoponymy (VIII): Meshchera, a Permian wedge between Volga Finns?”The ethnonym Meshchera [Мещёра] is not found in such very early Russian chronicles as Povest’ vremennyh let [“Nestor’s Chronicle”] (PSRL 1965), first appearing in