Open access Population structure in Argentina, by Muzzio et al., PLOS One (2018).
Abstract (emphasis mine):
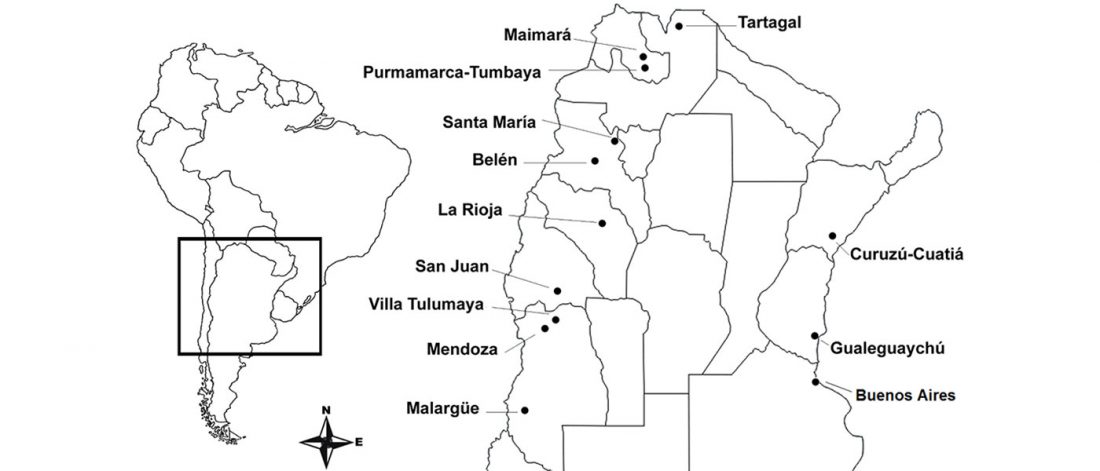
… Read the rest “Population structure in Argentina shows most European sources of South European origin”We analyzed 391 samples from 12 Argentinian populations from the Center-West, East and North-West regions with the Illumina Human Exome Beadchip v1.0 (HumanExome-12v1-A). We did Principal Components analysis to infer patterns of populational divergence and migrations. We identified proportions and patterns of European, African and Native American ancestry and found a correlation between distance to Buenos Aires and proportion of Native American ancestry, where the highest proportion corresponds to the Northernmost populations, which is also the furthest from the Argentinian capital. Most of